Antigen Density for Human and Murine Surface Markers

- On This Page
- Antigen Density for Human and Murine Surface Markers
On this page:
- Introduction
- ABC Bead Assay
- Experimental Protocol
- Standard Curve Creation
- Human Data
- Murine Data
- Fluorescent Bead Assay
- References
 Antigen Density of Common Human and Murine Markers
Antigen Density of Common Human and Murine Markers
Introduction
Tutorials for building multicolor flow cytometry panels always highlight the importance of antigen density - But why is it so important?
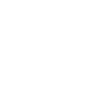
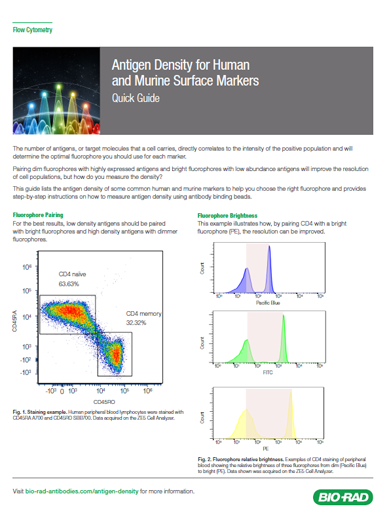
The number of antigens, or target molecules that a cell carries directly correlates to the intensity of the positive population and will determine the optimal fluorophore you should use for each marker. As a general rule you should pair bright fluorophores (e.g. PE) with low expressing markers and dimmer fluorophores (e.g. Pacific Blue) with highly expressed markers. Careful choice of fluorophores will help with the resolution of your cell populations. It is possible for bright fluorophores to be paired with highly expressed antigens, but better to avoid pairing dim fluorophores with low abundance antigens.
Care should be taken however, as fluorophore brightness can have both positive and negative effects.
A bright fluorophore will give better resolution, but have greater spread when compensated, and if placed on an abundant antigen, the increased spillover into adjacent channels may mask the signal from dimmer fluorophores.
 Fluorescent Spectraviewer
Fluorescent Spectraviewer
Use the Spectraviewer
 Build multicolor flow cytometry panels in just a few simple steps
Build multicolor flow cytometry panels in just a few simple steps
Go to Panel Builder
Fluorophore choice is relatively easy when only a few markers are required. However it becomes more complicated as you increase the number of fluorophores in your panels, especially if the antigen density is unknown or may change with cell subsets or activation state. Knowing the antigen density will help rank markers and pair them to similarly ranked fluorophores. The following guide provides advice on how to measure antigen density and lists the antigen density of some common markers.
If you know the antigen density for any given marker, you can input it in Bio-Rad’s panel builder tool, which will then automatically exclude sub-optimal fluorophores from your panel.
Antigen density can also be useful when determining the biological activity of ligands. The density of receptors has been shown to be related to the biological activity of ligands, e.g. cytokines (Gudipaty et al. 2001; Moraga et al. 2009; Reynes et al. 2000) and their subsequent efficacy.
Antibody Binding Capacity Bead Assays to Determine Antigen Density
There are several methods to measure antigen density, such as using the kinetics of antibody binding (Orlova et al. 2011) and radioactively labeled ligands. However the most common and simple method uses calibrated beads and flow cytometry.
The first type of bead assay relies on the binding of conjugated antibody to beads that have specific levels of antibody capture capacity via the Fc portion. The beads are labeled with the same antibody used to label the cells where antigen density is to be determined. Therefore any conjugated antibody can be used, providing the same batch with the same fluorophore:protein ratio (F:P ratio) is used to stain the beads and the cells in any one experiment.
The beads are individually stained at saturated levels, then combined and analyzed by flow cytometry to create a standard curve of the geometric mean fluorescence intensity plotted against the antibody binding capacity (ABC) of the beads. When cells are stained using the same antibody, antigen density is calculated by using the geometric mean to obtain an ABC value equal to the number of surface receptors, assuming a 1:1 binding ratio of antibody-to-surface receptor binding. We now go into further detail of how to obtain antigen density using these beads.
Antigen Density Protocol and Results
To determine antigen density on human and mouse cells, beads that bind either mouse or rat antibodies via the Fc portion were used. For mouse anti-human antibodies, Quantum Simply Cellular Mouse IgG Beads were used. All antibodies were titrated on human peripheral blood or murine splenocytes prior to starting the protocol to determine the optimal concentration for use. Saturating concentrations of antibody were also determined for the beads. An example of antibody titration and how the optimal concentration was chosen can be seen for Rat Anti-Mouse CD4 PE (MCA2691PE) in Figure 1. Visit our dedicated antibody titration page for more information on the benefits of titration.
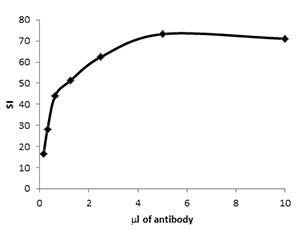
Fig. 1. Antibody titration. CD4 PE (MCA2691PE) was used to stain murine splenocytes at reducing concentrations. The stain index of the positive cells was plotted against antibody dilution (ml of antibody in 100 ml staining buffer) to reveal the optimal staining concentration.
After the optimal antibody concentrations were obtained by titration, human blood was Fc blocked using 10% human serum, and murine splenocytes were Fc blocked with 10% mouse serum. The samples were then stained with each antibody conjugated to PE. In parallel, the individual populations of beads, with increasing binding capacity, were stained with the same PE conjugated antibodies at saturating concentrations (also determined by titration), before being pooled ready for analysis by flow cytometry, following the guidelines stated on the datasheet. No Fc block was used for the beads. A population of beads was left unstained as a negative control. To ensure consistency within the experiment, for human and mouse samples, the staining was performed on the same samples, with the same protocol, for similar lengths of time, including the same number of wash steps using the same lot of antibody.
Beads and cells (human or mouse) were analyzed by flow cytometry immediately, ensuring the same instrument settings were used. This includes PMT voltages, laser power and compensation levels (if required). The forward and side scatter voltages were set to allow both cells and beads to be visible for ease of analysis (Figure 2). This was in addition to setting fluorescence detection to allow measurement of the negative and positive bead populations, and positive and negative cell populations. A low flow rate of 100-200 events per second is recommended and typically 1,000 to 1,500 events are collected for each bead population. We collected 30,000 cells for statistical significance and to ensure we could visualize rarer populations if required.
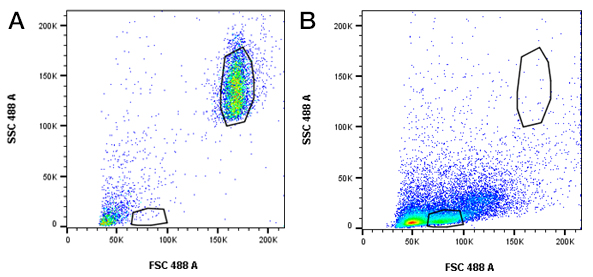
Fig. 2. Beads and cells are visible using the same forward and side scatter voltages. Forward and side scatter settings were used so that A, beads and B, splenocytes were visible and gates could be set without changing voltages.
Standard Curve Creation
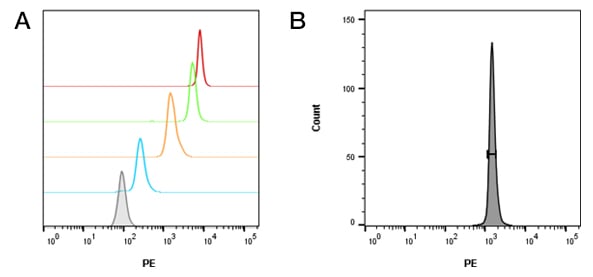
After acquisition of the beads for analysis, the individual populations of beads are gated using a half-height/full-width approach and the geometric mean for each bead population is recorded (Figure 3). The geometric mean value can be inputted into the QuickCal analysis template along with the ABC, as described on the datasheet, using the batch specific ID (Figure 4). This is important, as the antibody binding capacity is batch specific. Using these values, a linear regression curve can be plotted with the regression coefficient (r2) and detection threshold for each marker stained on cells for which an antigen density is required. R2 values as close to 1 are desired for accuracy and reproducibility of data.
C
Bead Population |
MFI (geo mean) |
ABC |
|---|---|---|
|
Blank |
90.9 |
|
|
1 |
258 |
7,311 |
|
2 |
1,520 |
44,263 |
|
3 |
4,760 |
161,876 |
|
4 |
7,249 |
269,561 |
Fig. 3. Bead staining and gating. A, populations of beads with increasing binding capacity show increased brightness (blue – low ABC, red – high ABC, grey – unstained). B, an example of half height/ full width gating on a bead population. C, table of values obtained from geometric means.
The geometric mean from the gated positive population from cells stained with titrated antibodies, minus the geometric mean from unstained cells (or from isotype control stained cells), can be read from the plotted standard curve to determine the antigen density on the cell surface, using the formula, y = mx +c (Figure 4). The Quick Cal program will perform this for you.
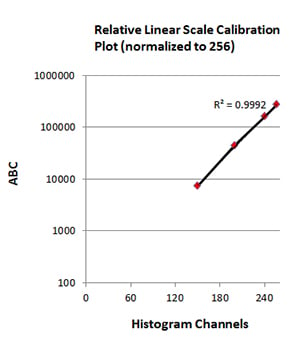
Fig. 4. Standard curve generation. The geometric mean fluorescence intensity values (histogram channels) were plotted against antibody binding capacity (ABC) to generate a standard curve. The R2 value was determined to assess accuracy and reproducibility.
Human Results
Peripheral blood from three donors was stained for 21 common surface markers conjugated to PE and acquired on the ZE5 Cell Analyzer (Figure 5) using the same conditions and instrument settings described earlier. Aside from some variation between donors, we can see the difference in surface expression levels, or antigen density of the markers, which allows us to choose an appropriate fluorophore when building a multicolor panel. The results shown here are from the ZE5 Cell Analyzer, other instruments may give slightly different results to those shown here.
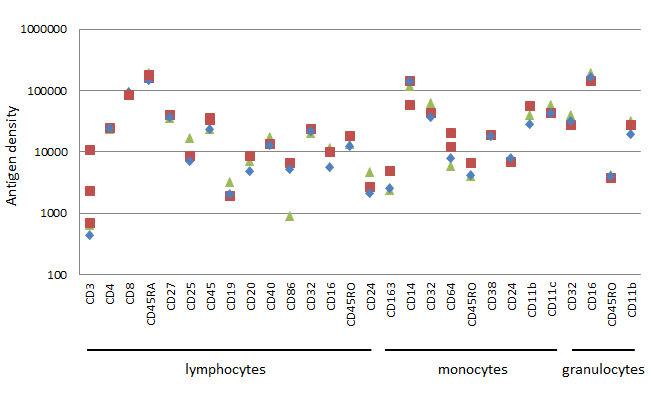
Fig. 5. Data from 21 common human markers. Geometric mean fluorescence intensity was obtained and converted into antigen density for each common surface marker and plotted on a log scale. The results for up to 4 independent experiments on normal human blood donors are shown.
Murine Results
Splenocytes from three different mice were stained for 16 common surface markers and acquired on the ZE5 Cell Analyzer (Figure 6) using the same conditions to collect stained beads as described earlier. This allowed us to calculate the surface antigen density of the 16 markers on fresh cells. From this data, we are able to recommend a bright fluorophore for CD25 and dim fluorophore for CD8.
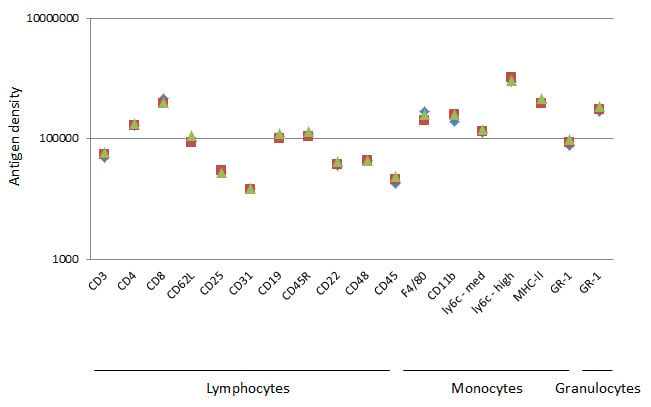
Fig. 6. Data from 16 common murine markers. Geometric mean fluorescence intensity was obtained and converted into antigen density for each common surface marker and plotted on a log scale. The results for 3 independent experiments on C57BL/6 mice are shown.
To optimize your data ensure that the number of cells positive for the marker of interest is sufficient, otherwise the detection threshold won’t be reached and you will be unable to determine an accurate antigen density. It is important to have high quality sample preparation to avoid high levels of dead cells and nonspecific binding, as this will influence the geometric mean fluorescence and therefore the antigen density.
The nature of the sample is also important and you need to ensure there is expression on the cell surface. This technique will allow you to determine and compare specific marker expression levels on different cell populations, as we have shown for GR-1 on monocytes and granulocytes. It also provides a comparison between activated and resting cells. However, to compare samples, the level of activation and the sample type would need to be the same. For example, do not compare fresh with frozen cells. As can be seen in Figure 7, the antigen density can vary between fresh and frozen cells for certain markers, such as CD19 and GR-1.
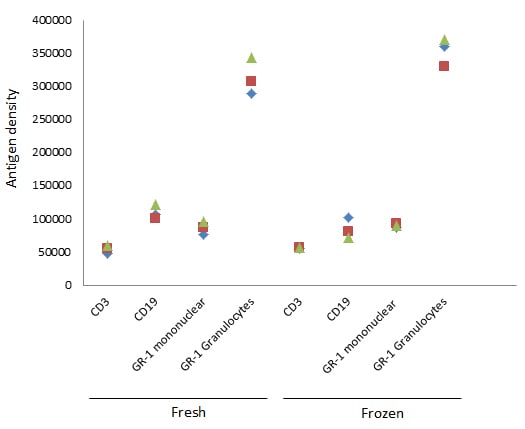
Fig. 7. Comparison between fresh and frozen cells. Comparing the antigen density of fresh and frozen murine splenocytes shows variation for some, but not all markers.
Fluorescent Bead Assays to Determine Antigen Density
The second type of bead assay relies on beads that are directly labeled with known amounts of a fluorophore, such as PE. Populations of beads with varying levels of PE (a negative and 3 or 4 varying positive populations) are combined and analyzed by flow cytometry to create a standard curve of the mean fluorescence intensity (MFI) plotted against the number of PE molecules. The number of PE molecules can be converted into antibodies per cell. When cells are stained with an antibody, the antigen density can be calculated from the MFI read against the standard curve, in a similar way to the antibody binding beads discussed above. This method also assumes that the degree of labeling is 1:1, which is true for PE but not necessarily the case for some fluorophore conjugated antibodies. Try Bio-Rad’s Quantum R-PE MESF Kits to measure antigen density, using beads of varying fluorescence intensity.
Bio-Rad has over 4,000 antibodies validated for flow cytometry, as well as helpful resources to guide your flow cytometry. These resources, in addition to listings of antibodies, kits, and controls can be accessed from our dedicated flow cytometry hub page or by clicking on the links below.
References
- Gudipaty et al. (2001). Regulation of P2X7 nucleotide receptor function in human monocytes by extracellular ions and receptor density. Am J Physiol Cell Physiol 280, 943-53.
- Moraga et al. (2009). Receptor density is key to the alpha2/beta interferon differential activities. Mol Cell Biol 17, 4778-87.
- Reynes et al. (2000). CD4+ T cell surface CCR5 density as a determining factor of virus load in persons infected with human immunodeficiency virus type 1. J Infect Dis 3, 927-32.
- Orlova et al. (2011). Distribution function approach to the study of the kinetics of IgM antibody binding to FcγRIIIb (CD16b) receptors on neutrophils by flow cytometry. J Theor Biol 290, 1-6.
Related flow cytometry products








