Mini-review: Understanding Innate Lymphoid Cells - A Diverse Immune Cell Population
Innate lymphoid cells (ILCs) are the most recently identified population of immune cells. They represent a distinct arm of the innate immune system and have key roles in inducing resistance to pathogens, regulation of autoimmune inflammation, metabolic homeostasis and tissue remodeling. Although they can be found in most tissues, they play important roles in barrier tissues such as the skin, lungs and intestine. This mini-review will outline the characteristics of ILCs as well as their biological identification and function. The development of ILCs and their role in the defense against pathogens and potential for immunotherapy are also addressed.
Overview of ILCs
Since the identification of leukocytes in the 19th century, several distinct lineages have been discovered, including myeloid and lymphocyte subsets (Addison 1843). Within the lymphocyte subgroup are B and T cells of adaptive immunity, and natural killer (NK) cells of the innate immune response. For most of the past 30 years, NK cells were the only known innate lymphoid cells (ILCs). However, at the end of the 20th century in 1997, lymphoid tissue inducer (LTi) cells were identified as a distinct subset of ILCs that promote the development of lymph nodes and Peyer’s patches during embryogenesis (Mebius et al. 1997). Groundbreaking research studies from 12 independent groups in 2008 and 2009 later identified a distinct group of non-T, non-B lymphocytes, generally classified as ILCs due to their similarities to NK cells (Spits and Di Santo 2011). ILCs represent a distinct arm of the innate immune system and can directly communicate with hematopoietic and non-hematopoietic cells to mediate immunity, inflammation and tissue homeostasis. They are also known to contribute to immune defense against pathogens, wound healing as well as maintenance of epithelial integrity.
Characterizing ILCs
The ILC family is characterized by a classic lymphoid morphology and the absence of rearranged antigen-specific receptors such that they do not exhibit antigen specificity, which is unique to T and B cells. The family includes conventional NK (cNK) cells, LTi cells and the recently identified non-cytotoxic group of ILCs, which are grouped as ILC1s, ILC2s and ILC3s. ILCs are separated into distinct groups based on phenotypic and functional traits, namely cytotoxic or killer ILCs (e.g. cNK cells) and non-cytotoxic helper-like ILCs (e.g. LTi cells) (Spits et al. 2013). The non-cytotoxic helper cells are separated into three groups: group 1 ILCs (ILC1s) consists of ILCs that secrete interferon-gamma (IFN-γ) and tumor necrosis factor (TNF); group 2 ILCs (ILC2s) produce type 2 cytokines such as interleukin (IL)-5 and IL-13 and group 3 ILCs (ILC3s) secrete IL-17 and IL-22, and includes LTi cells. The transcriptional profile, cytokine expression and effector functions of these non-cytotoxic ILCs strikingly resemble that of the CD4+ T helper cell lineage (Klose et al. 2014). It has also been proposed that NK cells are the innate form of CD8+ T cells (Hazenberg and Spits 2014).
ILCs are defined as cell lineage marker negative (Lin-) but express subunits of cytokine receptors including CD25 (IL-2 receptor-α) and CD127 (IL-7 receptor-α). The ILC1 and ILC3 groups are heterogeneous in both human and mouse, with two distinct subsets based on CD127 expression for ILC1s and natural cytotoxicity receptor (NCR) expression for ILC3s (Eberl et al. 2015). Table 1 outlines the unique characteristics of the various ILC subsets in human and mouse, including common markers for their identification, cytokine expression as well as their biological function.
Table 1. Characterization of ILC subsets in human and mouse
|
Marker Expression |
Cytokine/Chemokine Expression |
Cellular Activation |
Biological Function |
|---|---|---|---|---|
|
HUMAN |
||||
|
cNK cell
|
IFN-γ |
Activated by secretion of IL-15, IL-12 and IL-18 from DCs, neutrophils, monocytes and macrophages |
|
|
|
ieILC1 (CD127-) |
CD335 (NKp46) |
IFN-γ |
Activated by secretion of TGF-β, IL-12 and IL-18 from tissue-infiltrating monocytes |
|
|
ILC1 (CD127+) |
IFN-γ |
Activated by secretion of TGF-β, IL-12 and IL-18 from tissue-infiltrating monocytes |
|
|
|
ILC2 |
Areg |
Activated by IL-33, IL-25 and TSLP secreted by injured epithelial cells and possibly from a high calorie diet or IL-4 and PGD2 from basophils and mast cells |
Areg producing ILC2s
IL-5 and IL-13 producing ILC2s
|
|
|
ILC3 (NCR-) |
CCL3 |
Activated by CCL20 and IL-23 secretion from epithelial cells and DCs or IL-1β from macrophages via NLRP3 |
CCL3, LTs and IL-22 producing ILC3s
IFN-γ and IL-17 producing ILC3s
|
|
|
ILC3 (NCR+) |
CD127 |
CCL3 |
Activated by CCL20 and IL-23 secretion from epithelial cells and DCs or IL-1β from macrophages via NLRP3 |
CCL3, LTs and IL-22 producing ILC3s
IFN-γ and IL-17 producing ILC3s
|
|
MOUSE |
||||
|
cNK cell
|
IFN-γ |
Activated by secretion of IL-15, IL-12 and IL-18 from DCs, neutrophils, monocytes and macrophages |
|
|
|
ieILC1 (CD127-) |
IFN-γ |
Activated by secretion of TGF-β, IL-12 and IL-18 from tissue-infiltrating monocytes |
|
|
|
ILC1 (CD127+) |
CD127 |
IFN-γ |
Activated by secretion of TGF-β, IL-12 and IL-18 from tissue-infiltrating monocytes |
|
|
ILC2 |
CD127 |
Areg |
Activated by IL-33, IL-25 and TSLP secreted by injured epithelial cells and possibly from a high calorie diet or IL-4 and PGD2 from basophils and mast cells |
Areg producing ILC2s
IL-5 and IL-13 producing ILC2s
|
|
ILC3 (NCR-) |
CD127 |
CCL3 |
Activated by CCL20 and IL-23 secretion from epithelial cells and DCs or IL-1β from macrophages via NLRP3 |
CCL3, LTs and IL-22 producing ILC3s
IFN-γ and IL-17 producing ILC3s
|
|
ILC3 (NCR+) |
CD127 |
CCL3 |
Activated by CCL20 and IL-23 secretion from epithelial cells and DCs or IL-1β from macrophages via NLRP3 |
CCL3, LTs and IL-22 producing ILC3s
IFN-γ and IL-17 producing ILC3s
|
Adapted from Eberl et al. 2015. cNK, conventional natural killer cell; ieILC1, intraepithelial innate lymphoid cell group 1; TH1, T helper type 1; TH2, T helper type 2; Areg, amphiregulin; CCL3, chemokine (C-C motif) ligand 3; CCL20, chemokine (C-C motif) ligand 20; LTs, lymphotoxins; PGD2, prostaglandin D2; CRTH2, chemokine receptor homologous molecule expressed on TH2 lymphocytes; TSLP, thymic stromal lymphopoietin; VLA1, very late antigen integrin 1; VLA2, very late antigen integrin 2; NLRP3, nucleotide-binding domain, leucine-rich-containing family, pyrin domain-containing-3; DC, dendritic cells; LNs, lymph nodes; NCR, natural cytotoxicity receptor.
Development of ILCs
The current understanding of how ILCs develop has been determined through the use of relevant mouse models. Analogous human precursors to those identified in mice have not yet been clearly defined; however, we highlight the current findings on human ILC development.
ILC development in mice
To elucidate the developmental process of ILCs in mice, scientists first demonstrated that ILCs and adaptive immune lymphocytes originate from a common lymphoid progenitor (CLP) (Klose et al. 2014, Cherrier et al. 2012). Two populations of cells have been shown to be downstream of CLPs and demonstrate the ability to generate all ILC populations but cannot differentiate into B and T cells. These are α-lymphoid progenitors (α-LPs) and early innate lymphoid progenitors (EILPs). Both of these cells are phenotypically similar to CLPs but lack specific transcription factors and markers necessary for B and T cell development. The α-LP population differs from CLPs in their expression of the integrin α4β7, CXCR6 and the absence of CD135 (Flt3) (Zook and Kee 2016). Similarly, the transcriptome of EILPs is similar to CLPs except that EILPs do not express ILC-lineage specific transcription factors such as RORγt, GATA-3, T-bet or Eomes.
Downstream of EILPs or α-LPs, two distinct ILC lineages have been identified. These are the killer ILC (NK cells) and the various helper cytokine producing ILC subsets (ILC1s, ILC2s and ILC3s). A key distinction between the two lineages is based on the dependency on the transcription factor GATA-3. Helper ILC subsets require GATA-3 for differentiation, whereas NK cells can develop normally in GATA-3-deficient mice (Diefenbach et al. 2014). The developmental pathway for NK cells from a specified NK progenitor to a mature NK cell is understood in greater detail than helper ILC development, partly because their existence has been known for longer. A revised NK progenitor (rNKP) has recently been described that shares a similar phenotype to the CLP but lack expression of CD135 and Id2, and express CD122 and CD127 (Fathman et al. 2011). Notably, cNK cells upregulate Id2 after lineage specification (Yokota et al. 1999). These rNKPs demonstrate significant commitment to NK cell differentiation both in vitro and in vivo. On their way to becoming mature NK cells (mNKs), rNKPs have been shown to first pass through an immature NK cell (iNK) stage. iNKs express NK1.1 but do not yet express CD49 (DX5) (Zook and Kee 2016), and their phenotype is similar to ILC1s. However, iNK cells are distinguished from ILC1s via the expression of CD127, the integrin α4β7, and CD49a (also known as DX5a or α1). The iNK cell stage marks the onset of the expression of Ly49 receptors, which are necessary for mNK function. iNK cells undergo a maturation process to become mNKs, however not all mNKs are functionally equivalent (Zook and Kee 2016). The maturation process is measured by differential expression of the costimulatory receptor CD27 and the integrin CD11b (also known as αM). The cytotoxic capacity of developing NKs improves with maturation and the upregulation of CD11b expression (Bezman et al. 2012).
To determine a common progenitor for helper like ILCs, Id2 reporter mice (Rawlins et al. 2009) were used to identify Id2+ lymphoid cells in the bone marrow and fetal liver of adult mice (Cherrier et al. 2012). These cells were characterized by the expression of CD127 and intermediate levels of Sca-1 and c-kit. These cells were found to be Lin- Id2+ CD127+ α4β7+ CD25- CD135-, and are referred to as common helper-like ILC progenitor (CHILP). CHILPs do not express the transcription factors generally associated with ILCs (e.g. RORγt, Eomes and T bet), indicating that they are not committed to a specific ILC lineage (Klose et al. 2014). Transfer of CHILPs into alymphoid mice demonstrate their capacity to differentiate into both subsets of ILC1s and ILC3s as well as ILC2s, but not cNKs (Diefenbach et al. 2014). This suggests that all cytokine producing helper ILCs originate from a common progenitor (CHILP) and that cNKs branch off from EILPs. Two factors essential for the generation of CHILPs are the transcription factors NFIL3 and EST1, which act at the EILP or α-LP stage (Zook and Kee 2016).
The CHILP population of cells has been shown to be heterogeneous, with a subset of cells expressing the transcription factor PLZF (Constantinides et al. 2014). PLZF+ CHILPs express GATA-3 and lack the capacity to generate the LTi cell subset of ILC3s. These cells are referred to as helper ILC progenitors (ILCPs) (Zook and Kee 2016). ILCPs can differentiate into ILC1s, ILC2s and ILC3s. However, the expression of PLZF is specifically required for the development of ILC2s and liver ILC1s (Constantinides et al. 2014).
The discussed findings represent only a portion of the developmental program of ILCs (summarized in figure 1), and several questions remain such as whether STAT transcription factors play a role in their development as is observed in T cell development. Identifying the intermediates in these developmental programs will provide significant insight into how to control ILCs in disease or exploit them for therapeutic purposes as discussed in section 4 (Zook and Kee 2016).
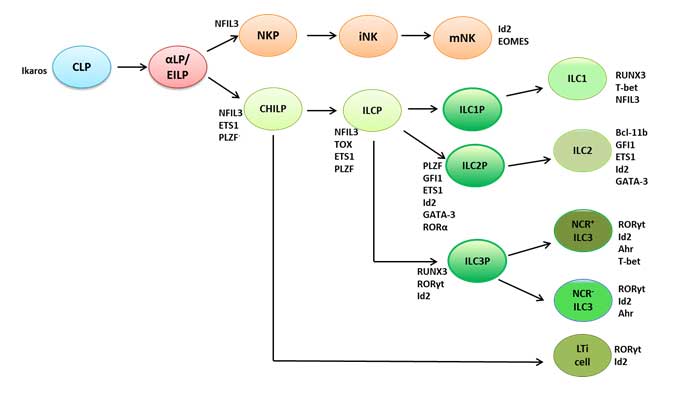
Figure 1. Development of mouse ILCs and the transcription factors involved at each developmental stage
Adapted from Zook and Kee 2016. CLP, common lymphoid progenitor; α-LP, α-lymphoid progenitors; EILP, early innate lymphoid progenitors; NKP, natural killer cell progenitor; iNK, immature natural killer cell; mNK, mature natural killer cell; CHILP, common helper-like ILC progenitor; ILCP, helper ILC progenitors; ILC1P, innate lymphoid cell group 1 progenitor; ILC2P, innate lymphoid cell group 2 progenitor; ILC3P, innate lymphoid cell group 3 progenitor; ILC1, innate lymphoid cell group 1; ILC2, innate lymphoid cell group 2; ILC3, innate lymphoid cell group 3; NCR, natural cytotoxicity receptor; LTi, lymphoid tissue inducer, NFIL3, nuclear factor interleukin-3 regulated protein; Id2, inhibitor of DNA binding 2; EOMES, eomesodermin; ETS1, ETS proto-oncogene 1, transcription factor; PLZF, promyelocytic leukemia zinc finger protein; TOX, thymocyte selection-associated HMG-box; GFI1, growth factor independent 1 transcription repressor; GATA-3, GATA binding protein 3; RORα, retinoic acid receptor-related orphan receptor alpha, RUNX3, Runt related transcription factor 3; RORγt, retinoic acid receptor-related orphan receptor gamma in thymus; Bcl-11b, B cell CLL/lymphoma 11B; Ahr, aryl-hydrocarbon receptor.
ILC development in humans
The developmental pathways of ILCs in humans as well as the transcription factors that regulate the process are less well defined compared to ILC development in mice. A recent study identified a progenitor population in human secondary lymphoid tissues that expressed the transcription factor RORγt and was capable of generating all known ILC subsets, including NK cells, but not other leukocyte populations (Scoville et al. 2016). Mouse studies have demonstrated that only ILC3s express RORγt. In contrast, these recent findings demonstrate that the human precursor cell, human peripheral blood NK cells and all mature ILC populations express RORγt (Scoville et al. 2016). Therefore, the research to date suggests that all human ILCs can be generated via a RORγt+ developmental pathway from a common progenitor in secondary lymphoid organs.
Human NK cell development has been described in further detail. The current model based on earlier studies, suggests a four-stage development process based on the expression of CD34, CD117 (cKit), CD94 and CD56 (Hazenberg and Spits 2014). Cells in the first stage are CD34+ CD56-, and have been suggested to overlap with the human common lymphoid progenitor. Stage 2 cells are CD34+ CD117+. Stage 3 cells are described as iNK cells, and are CD117+ CD56+. Cells in the final stage are considered mature NK cells and are CD56+ CD94+ CD16-. Other mature NK cells that are CD56+ CD94+ CD16+ have also been identified and are described as belonging to stage 5 (Hazenberg and Spits 2014).
Figure 2 provides a summary of the current knowledge of human ILC development.
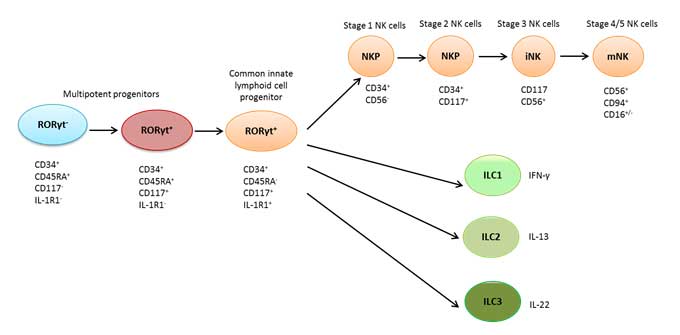
Figure 2. Model of human ILC development in secondary lymphoid tissues
Adapted from Scoville et al. 2016 and Hazenberg and Spits 2014. ILC1, innate lymphoid cell group 1; ILC2, innate lymphoid cell group 2; ILC3, innate lymphoid cell group 3; NKP, natural killer cell progenitor; iNK, immature natural killer cell; mNK, mature natural killer cell; RORγt, retinoic acid receptor-related orphan receptor gamma in thymus.
Function of ILCs in pathogen defense
Despite being the most recent immune cells to be identified, ILCs have already been shown to be key players in several aspects of the immune response. Specifically, they are necessary for inducing resistance to pathogens, regulation of autoimmune inflammation, metabolic homeostasis and tissue remodeling. Their role in defense against pathogens was anticipated given their prompt activation and early response to infection. In addition, they were initially identified at barrier surfaces, suggesting their importance in host protection and homeostasis (Klose and Artis 2016).
All ILC subsets have been implicated in inflammatory diseases using a variety of models. Mouse models of infection show that ILCs are particularly adept at controlling pathogens that use the mucosa to penetrate the host (Klose and Artis 2016). However, very little is known about human ILCs in host defense at barrier surfaces. Further research is needed to fully understand how human ILCs function in host protection. Here we focus on studies using relevant mouse models and provide insight into human studies where applicable.
NK cells and ILC1s in pathogen defense
T-bet-deficient mice, which lack ILC1s but have normal NK cell numbers, have been used to study the role of ILC1s in infectious diseases. It has been shown that CD127+ intestinal ILC1s play a role in protection against Toxoplasma gondii infection since T-bet deficient mice succumb to T. gondii (Klose et al. 2014). Because T-bet deficient NK cells are incapable of migration to infection sites, NK cells may also play a key role in controlling T. gondii infection (Jenne et al. 2009). T-bet dependent ILC1s are also necessary for protecting epithelial barriers from Salmonella enterica infection (Klose et al. 2013). Ragγc-/- mice which lack ILCs as well as T and B cells were used to demonstrate a critical role for ILC1s in host defense against acute Clostridium difficile (Abt et al. 2015). This study demonstrated that loss of T-bet expressing ILC1s increased susceptibility to C. difficile. ILC1s and NK cells have also been implicated in protection against colitis using Tbx21-/-Rag2-/- mice which lack T-bet ILC1s and NK cells (Powell et al. 2012). In humans, an increased number of ILC1s was observed in the inflamed tissues of patients with various autoimmune diseases, such as Crohn’s disease (Bernink et al. 2013). These findings demonstrate the importance of ILC1s in protection against various pathogens as well as in autoimmunity and inflammation.
ILC2s in pathogen defense
Long before the discovery of ILC2s, potent Th2-like responses to infection were observed in T cell-deficient mice (Fort et al. 2001). ILC2s demonstrate a similar cytokine profile as Th2 T cells and have been shown to be key mediators of the immune response to various pathogens (Moro et al. 2010). They are particularly known for mediating resistance to helminths such as Nippostrongylus brasiliensis, Strongyloides venezuelensis and Trichuris muris (Fallon et al. 2006, Zaiss et al. 2006, Yasuda et al. 2012). Upon activation by IL-25 and IL-33, secreted by tuft cells in the intestines, ILC2s secrete IL-4, IL-5, IL-13 and amphiregulin to combat helminth infection (Klose and Artis 2016). IL-5 is necessary for activating eosinophil function and amphiregulin initiates the repair of epithelial cells. IL-13 plays a key role in removing helminths by inducing mucus production, smooth muscle contraction, eotaxin release and recruitment of M2 macrophages (Artis and Spits 2015, Yasuda et al. 2012). In addition to their effect on helminths, amphiregulin secretion by ILC2s has also been shown to play an important role in promoting lung tissue homeostasis after influenza virus infection (Monticelli et al. 2011).
ILC3s in pathogen defense
ILC3s have been shown to be critical for protection against intestinal pathogens such as Salmonella typhimurium, Citrobacter rodentium, Candida albicans and rotaviruses (Klose and Artis 2016). The main mechanism of action involves ILC3 secretion of IL-22. IL-22 mediates resistance to these pathogens by binding to a heterodimeric receptor composed of both the IL-22 receptor α1 and IL-10 receptor β (IL-22Rα1-IL-10Rβ) on epithelial cells. This triggers activation of a signaling cascade leading to STAT-3 phosphorylation, cell proliferation, secretion of antimicrobial peptides such as RegIIIβ, RegIIIγ, S108a and S109a as well as fucosylation of epithelial cells (Wolk et al. 2004, Zheng et al. 2008). It is this fucosylation of the intestinal epithelium that induces protection against S. typhimurium infection (Goto et al. 2014). IL-22 secretion by ILCs has also been shown to mediate STAT-1 dependent resistance to rotaviruses. Upon infection with rotaviruses, IL-1α is released from epithelial cells, which stimulates IL-22 production by ILC3s leading to STAT-1 activation (Klose and Artis 2016). ILC3 mediated secretion of IL-22 also mediates protection against C. rodentium, a model organism for attaching and effacing Escherichia coli (Zheng et al. 2008).
It should be noted that the mouse models used to study the role of ILC3s in the response to pathogens demonstrate certain flaws that could confound the findings since they were designed to assess the role of ILC3s independently of the adaptive immune response. For example, studies using RAG-deficient mice, which lack T and B cells, compared to Rag-/-Il2rg-/- mice, which lack T and B cells as well as ILCs show an important role for ILC3s in defense against C. rodentium (Klose and Artis 2016). However, these models do not factor in the possibility that ILC3s and T cells could have redundant functions, and T cells could compensate for ILC3 roles in pathogen defense.
To address this concern, another approach has been to treat mice with anti-CD90 antibody that depletes most ILCs but retains T cell and B cell function. Using this system, ILC3s were shown to be necessary for controlling symbiotic bacteria such as the Alcaligenes species (Cording et al. 2016). However, more specific systems for studying ILC3 function in pathogen defense have been generated. An Nkp46Cre/+ system that interferes with the development and function of NCR+ ILC3s was used to determine the role of this subset of ILC3s during C. rodentium infection (Song et al. 2015, Rankin et al. 2016). It was shown that NCR+ ILC3s are dispensable in the presence of T cells for controlling C. rodentium, indicating the importance of the mouse model used to study the role of ILCs in infection. It also demonstrates the complementary and redundant functions of innate and adaptive lymphocytes.
Studies in humans demonstrate that RORγ-deficient patients, who lack TH17 and ILC3s, are more susceptible to C. albicans infection (Okada et al. 2015). Similar results were observed in RORγt-deficient-mice in that these mice also failed to control C. albicans infection (Gladiator et al. 2013). Notably, resistance to C. albicans is not T cell dependent, as RAG-deficient mice can control C. albicans. However, RAG-deficient mice in which ILCs have been depleted or those in which IL-17 or IL-23 (key ILC3 effector cytokines) have been neutralized, are susceptible to C. albicans (Gladiator et al. 2013). These studies clearly implicate ILC3s as important mediators in the defense against C. albicans in both humans and mice.
Taken together, ILCs have been shown to be promising targets for boosting immunity to a number of pathogens, particularly those that infect mucosal surfaces.
ILCs in disease and immunotherapy
Our current understanding of ILCs indicates that they are key immune mediators that play a vital role in early immune responses. It is now clearly demonstrated that ILCs respond to signals from infected or injured tissues, produced by macrophages, dendritic cells or stromal cells (Table 1), similar to T cells. Accordingly, ILC1s, ILC2s and ILC3s are considered the innate counterparts of TH1, TH2 and TH17 subsets of T helper cells, respectively. Whereas activation of naïve T cells requires several days, ILCs are promptly activated to produce effector cytokines that were previously thought to be limited to T cells (Cording et al. 2016). Therefore, ILCs represent novel therapeutic targets for diseases where preventive and therapeutic strategies could not be developed by targeting T cells. Below we outline the role of ILCs in allergy, autoimmunity and cancer. We also discuss ILC-based therapeutic approaches for addressing these diseases in which ILCs have critical functions.
ILCs in allergy and inflammation
The cytokines that mediate ILC2 function such as IL-33, IL-5, IL-4 and IL-13 have all been shown to be associated with susceptibility to atopic diseases such as allergy and allergic rhinitis (Li et al. 2015, Klose and Artis 2016). This indicates a role for ILC2s in the pathology of these diseases. In fact, mouse studies demonstrate that ILC2s mediate detrimental responses in other atopic diseases such as asthma, atopic dermatitis and chronic rhinosinusitis (Kim et al. 2013, Halim et al. 2012, Imai et al. 2013). Furthermore, reconstitution of ILC-deficient mice with ILC2s induced asthma like symptoms (Halim et al. 2012). Patients with atopic dermatitis or chronic rhinosinusitis also show activated ILC2s in skin lesions or nasal polyps, respectively (Salimi et al. 2013, Mjösberg et al. 2011). Moreover, IL-4 secreted by basophils can stimulate ILC2s to promote inflammation in the lungs and skin (Motomura et al. 2014, Kim BS et al. 2014). These data based on genome wide patient studies and functional analyses in mice indicate that ILC2s are involved in the pathogenesis of atopic (hyperallergic) diseases (Klose and Artis 2016).
While ILC2s are predominantly involved in allergic responses, detrimental ILC3 responses have been demonstrated in inflammatory diseases. Mouse models of colitis induced by Helicobacter hepaticus, Helicobacter typhlonius, S typhimurium or anti-CD40 all indicate a role for ILC3s in the pathogenesis of this disease (Klose and Artis 2016). In patients with Crohn’s disease, impaired number and function of ILC3s has been observed (Geremia et al. 2011, Fuchs et al. 2013). The studies demonstrated increased IL-17 production by ILC3s in patients with Crohn’s disease compared to patients without inflammatory bowel disease (IBD).
Although ILC2s are important for lung inflammation, ILC3s have been shown to regulate airway hyper-reactivity induced by a high-fat diet (Kim HY et al. 2014). IL-17 deficient or RAG-deficient mice depleted of ILCs via anti-CD90 administration have been utilized to show that IL-17 derived from ILC3s regulates airway hyper-reactivity. This was shown to be mediated by IL-1β released via activation of the nucleotide-binding domain, leucine-rich-containing family, pyrin domain-containing-3 (NLRP3) inflammasome (Kim HY et al. 2014). Similar to ILC2s, a role for ILC3s in skin inflammation has also been suggested since patients with psoriasis vulgaris demonstrate an accumulation of ILC3s (Teunissen et al. 2014). A mouse model of psoriasis confirmed the contribution of ILC3s to the disease phenotype (Pantelyushin et al. 2012).
The importance of ILCs in chronic inflammation demonstrates their potential for therapeutic targeting as discussed below.
ILCs in cancer
The role of NK cells in cancer immune surveillance and in mediating antitumor responses has been well documented (Morvan and Lanier 2016). However, these studies were conducted long before the discovery of ILC1s. It is therefore likely that some of the functions attributed to NK cells could be mediated by ILC1s as phenotypically distinguishing these two cell types without the use of reporter mice is difficult (Spits et al. 2016). Moreover, there is currently no definitive strategy to selectively deplete NK cells and ILC1s (Spits et al. 2016). However, a role for ILC1s in cancer was suggested by the identification of an ILC1 or ILC1-like population with similarities to salivary gland ILC1s or NK cells, which were shown to be involved in the surveillance of certain oncogene-induced breast and prostate mouse tumors (Dadi et al. 2016).
ILC3s have been shown to induce anti-cancer as well as cancer-promoting effects; however, a role for ILC2s in cancer has been suggested but has not been formally demonstrated (Klose and Artis 2016). Studies using several strains of knockout mice demonstrated that ILC3s are required for tumor rejection. Using a transplantable tumor model of IL-12 expressing B16 melanoma cells, intratumoral injection of ILC3s was shown to inhibit tumor growth and induce changes in the tumor microvasculature in an IL-12 dependent manner (Eisenring et al. 2010). Furthermore, NCR+ ILC3s were found in human non-small-cell lung cancer, and they were speculated to promote the formation of protective tertiary lymphoid structures (Carrega et al. 2015). Secretion of IL-22 by ILC3s is also implicated in colitis-associated cancer. CCR6+ ILC3s were shown to be the main source of IL-22 in a mouse model of colitis-associated cancer, and depletion of ILCs or neutralization of IL-22 resulted in reduced tumor burden (Kirchberger et al. 2013).
Although the described studies clearly demonstrate a role for ILCs in cancer, further research is needed to fully understand the mechanisms by which ILCs interact with tumor cells, particularly in humans.
ILCs in immunotherapy
Because ILCs function upstream of the developing immune response to infection or inflammation, as discussed in previous sections of this mini-review, they represent the opportunity for novel immunotherapeutic strategies. Within the context of pathogen defense, approaches aimed at activating ILCs early in the immune response to infection could lead to substantial improvements to immunotherapy against pathogens. In contrast, late activation of ILCs could also reinforce the adaptive response leading to improved outcomes towards pathogen elimination (Cording et al. 2016). The limitation of such an approach however is that no specific activators of the distinct ILC subsets have been identified, since the cytokines that stimulate their function also induce T cells. However, local application of these cytokines in a time dependent manner could alleviate the potential risk of systemic activation. Another approach to treating pathogen mediated diseases through targeting of ILCs is to develop more efficient vaccines that incorporate antigens as well as adjuvants that activate the relevant ILC subsets.
In addition to treating pathogen induced diseases, strategies to activate ILC2s in particular could significantly impact the treatment of diseases such as obesity and diabetes. ILC2s can block pathogenic type 3 mediated responses in white adipose tissue, thus preventing its accumulation which causes obesity (Brestoff et al. 2015). ILC2s also block the increase in blood sugar, thereby inducing beneficial effects for the treatment of diabetes (Cording et al. 2016). ILC2 activation may also aid in the regulation of IBD and autoimmunity mediated by type 3 responses such as arthritis and multiple sclerosis (Klose and Artis 2016). Activation of ILC3s may also lead to inhibition of allergic inflammation induced by type 2 responses, and could prevent tissue invasion by microbiota via secretion of IL-17 and IL-22 (Lochner et al. 2010).
On the other hand, inhibition of ILC activity could also be applied to counter certain immunopathology in which ILCs induce detrimental effects. For instance, the use of neutralizing antibodies to inhibit the effector functions of ILC1s and ILC3s in IBD and pathogen induced colitis, or ILC2s in allergy and fibrosis, could successfully treat these diseases (Cording et al. 2016).In this case, however, a more targeted approach to inhibiting these ILC functions should be applied to avoid systemic deregulation of immunity. Timing of the administration of neutralizing antibodies should also be considered as ILCs also play key roles in maintaining homeostasis (Cording et al. 2016).
Manipulating ILCs for clinical use would be advantageous for combating devastating disease. However further research is needed to determine how to specifically target these cells without exacerbating or eliciting immunopathology.
References
- Abt MC et al. (2015). Innate immune defenses mediated by two ILC subsets are critical for protection against acute Clostridium difficile infection. Cell Host Microbe 18, 27-37.
- Addison W (1843). Experimental and Practical Researches on the Blood: Second Series. Prov Med J Retrosp Med Sci 6, 444-445.
- Artis D and Spits H (2015). The biology of innate lymphoid cells. Nature 517, 293-301.
- Bernink JH et al. (2013). Human type 1 innate lymphoid cells accumulate in inflamed mucosal tissues. Nat Immunol 14, 221-229.
- Bezman NA et al. (2012). Immunological Genome Project Consortium. Molecular definition of the identity and activation of natural killer cells. Nat Immunol 13, 1000-1009.
- Brestoff JR et al. (2015). Group 2 innate lymphoid cells promote beiging of adipose and limit obesity. Nature 519, 242-246.
- Carrega P et al. (2015). NCR+ ILC3 concentrate in human lung cancer and associate with intratumoral lymphoid structures. Nat Commun 6, 8280.
- Cherrier M et al. (2012). Notch, Id2, and RORγT sequentially orchestrate the fetal development of lymphoid tissue inducer cells. J Exp Med 209, 729-740.
- Constantinides MG et al. (2014). A committed precursor to innate lymphoid cells. Nature 508, 397-401.
- Cording S et al. (2016). Innate lymphoid cells in defense, immunopathology and immunotherapy. Nat Immunol 17, 755-757.
- Dadi S et al. (2016). Cancer immunosurveillance by tissue-resident innate lymphoid cells and innate-like T cells. Cell 164, 365-377.
- Diefenbach A et al. (2014). Development, differentiation and diversity of innate lymphoid cells. Immunity 41, 354-365.
- Eberl G et al. (2015). The brave new world of innate lymphoid cells. Nat Immunol 16, 1-5.
- Eisenring M et al. (2010). IL-12 initiates tumor rejection via lymphoid tissue-inducer cells bearing the natural cytotoxicity receptor NKp46. Nat Immunol 11, 1030-1038.
- Fallon PG et al. (2006). Identification of an interleukin (IL)-25-dependent cell population that provides IL-4, IL-5, and IL-13 at the onset of helminth expulsion. J Exp Med 203, 1105-1116.
- Fathman JW et al. (2011). Identification of the earliest natural killer cell-committed progenitor in human bone marrow. Blood 118, 5439-5447.
- Fort MM et al. (2001). IL-25 induces IL-4, IL-5, and IL-13 and Th2 associated pathologies in vivo. Immunity 15, 985-995.
- Fuchs A et al. (2013). Intraepithelial type 1 innate lymphoid cells are a unique subset of IL-12- and IL-15-responsive IFN-γ-producing cells. Immunity 38, 769-781.
- Geremia A et al. (2011). IL-23-responsive innate lymphoid cells are increased in inflammatory bowel disease. J Exp Med 208, 1127-1133.
- Gladiator A et al. (2013). Cutting edge: IL-17-secreting innate lymphoid cells are essential for host defense against fungal infection. J Immunol 190, 521-525.
- Goto Y et al. (2014). Innate lymphoid cells regulate intestinal epithelial cell glycosylation. Science 345, 1254009.
- Halim TY (2012). Lung natural helper cells are a critical source of Th2 cell-type cytokines in protease allergen-induced airway inflammation. Immunity 36, 451-463.
- Hazenberg MD and Spits H (2014). Human innate lymphoid cells. Blood 124, 700-707.
- Imai Y et al. (2013). Skin-specific expression of IL-33 activates group 2 innate lymphoid cells and elicits atopic dermatitis-like inflammation in mice. Proc Natl Acad Sci USA 110, 13921-13926.
- Jenne CN et al. (2009). T-bet-dependent S1P5 expression in NK cells promotes egress from lymph nodes and bone marrow. J Exp Med 206, 2469-2481.
- Kim BS et al. (2013). TSLP elicits IL-33-independent innate lymphoid cell responses to promote skin inflammation. Sci Transl Med 5, 170ra16.
- Kim BS et al. (2014). Basophils promote innate lymphoid cell responses in inflamed skin. J Immunol 193, 3717-3725.
- Kim HY et al. (2014). IL-17 producing innate lymphoid cells and the NLRP3 inflammasome facilitate obesity-associated airway hyperreactivity. Nat Med 20, 54-61.
- Kirchberger S et al. (2013). Innate lymphoid cells sustain colon cancer through production of interleukin-22 in a mouse model. J Exp Med 210, 917-931.
- Klose CS and Artis D (2016). Innate lymphoid cells as regulators of immunity, inflammation and tissue homeostasis. Nat Immunol 17, 765-774.
- Klose CS et al. (2014). Differentiation of type 1 ILCs from a common progenitor to all helper-like innate lymphoid lineages. Cell 157, 340-356.
- Klose CS et al. (2013). A T-bet gradient controls the fate and function of CCR6-RORγt+ innate lymphoid cells. Nature 494, 261-265.
- Li J et al. (2015). Discovering susceptibility genes for allergic rhinitis and allergy using a genome-wide association study strategy. Curr Opin Allergy Clin Immunol 15, 33-40.
- Lochner M et al. (2010). Microbiota-induced tertiary lymphoid tissues aggravate inflammatory disease in the absence of RORγt and LTi cells. J Exp Med 208, 125-134.
- Mebius RE et al. (1997). Developing lymph nodes collect CD4+CD3-LTβ+ cells that can differentiate to APC, NK cells, and follicular cells but not T or B cells. Immunity 7, 493-504.
- Mjösberg JM et al. (2011). Human IL-25- and IL-33-responsive type 2 innate lymphoid cells are defined by expression of CRTH2 and CD161. Nat Immunol 12, 1055-1062.
- Montaldo E et al. (2016). Human innate lymphoid cells. Immunol Lett doi: 10.1016/j.imlet.2016.01.007. [Epub ahead of print]
- Monticelli LA et al. (2011). Innate lymphoid cells promote lung tissue homeostasis following acute influenza virus infection. Nat Immunol 12, 1045-1054.
- Moro K et al. (2010). Innate production of TH2 cytokines by adipose tissue-associated c-Kit+Sca-1+ lymphoid cells. Nature 463, 540-544.
- Morvan MG and Lanier LL (2016). NK cells and cancer: you can teach innate cells new tricks. Nat Rev Cancer 16, 7-19.
- Motomura Y et al. (2014). Basophil-derived interleukin-4 controls the function of natural helper cells, a member of ILC2s, in lung inflammation. Immunity 40, 758-771.
- Okada S et al. (2015). Impairment of immunity to Candida and Mycobacterium in humans with bi-allelic RORC mutations. Science 349, 606-613.
- Pantelyushin S et al. (2012). Rorγt+ innate lymphocytes and γδ T cells initiate psoriasiform plaque formation in mice. J Clin Invest 122, 2252-2256.
- Powell N et al. (2012). The transcription factor T-bet regulates intestinal inflammation mediated by interleukin-7 receptor+ innate lymphoid cells. Immunity 37, 674-684.
- Rankin LC et al. (2016). Complementarity and redundancy of IL-22 producing innate lymphoid cells. Nat Immunol 17, 179-186.
- Rawlins EL et al. (2009). The Id2+ distal tip lung epithelium contains individual multipotent embryonic progenitor cells. Development 136, 3741-3745.
- Salimi M et al. (2013). A role for IL-25 and IL-33 driven type-2 innate lymphoid cells in atopic dermatitis. J Exp Med 210, 2939-2950.
- Scoville SD et al. (2016). A progenitor cell expressing transcription factor RORγt generates all human innate lymphoid cell subsets. Immunity 44, 1140-1150.
- Song C et al. (2015). Unique and redundant functions of Nkp46+ ILC3s in models of intestinal inflammation. J Exp Med 212, 1869-1882.
- Spits H and Di Santo JP (2011). The expanding family of innate lymphoid cells: regulators and effectors of immunity in tissue remodeling. Nat Immunol 12, 21-27.
- Spits H et al. (2013). Innate lymphoid cells — a proposal for uniform nomenclature. Nat Rev Immunol 13, 145-149.
- Spits H et al. (2016). NK cells and type 1 innate lymphoid cells: partners in host defense. Nat Immunol 17, 758-764.
- Teunissen MBM et al. (2014). Composition of innate lymphoid cell subsets in the human skin: enrichment of NCR+ ILC3 in lesional skin and blood of psoriasis patients. J Invest Dermatol 134, 2351-2360.
- Wolk K et al. (2004). IL-22 increases the innate immunity of tissues. Immunity 21, 241-254.
- Yasuda K et al. (2012). Contribution of IL-33-activated type II innate lymphoid cells to pulmonary eosinophilia in intestinal nematode-infected mice. Proc Natl Acad Sci USA 109, 3451-3456.
- Yokota Y et al. (1999). Development of peripheral lymphoid organs and natural killer cells depends on the helix-loop-helix inhibitor Id2. Nature 397, 702-706.
- Zaiss DM et al. (2006). Amphiregulin, a TH2 cytokine enhancing resistance to nematodes. Science 314, 1746.
- Zheng Y et al. (2008). Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat Med 14, 282-289.
- Zook EC and Kee BL (2016). Development of innate lymphoid cells. Nat Immunol 17, 775-782.








