Mini-review: An overview of T cell receptors
The immune system has a near limitless capacity for detecting abnormalities. This remarkable ability for selfinterrogation is achieved by the related structures of two molecules, immunoglobulins and T cell receptors (TCR). The TCR, a defining structure of T cells, is a transmembrane heterodimer consisting of either an alpha and beta chain or delta and gamma chain linked by a disulphide bond. Within these chains are complementary determining regions (CDRs) which determine the antigen to which the TCR will bind. TCRs activate the T cells in which they reside leading to a plethora of immune responses. Harnessing the power of this response and of TCR specificity is leading to a new generation of extremely promising immunotherapies.
Overview of TCRs
Initial descriptions of TCRs were made 30 years ago primarily through similarity with immunoglobulin DNA sequences. From this beginning, a clearer picture of TCRs as a pair of clone-specific, heterodimeric polypeptide chains consisting of both constant and variable regions has developed (Clambey et al. 2014). In humans, the majority of T cells express a TCR composed of alpha (α) and beta (β) chains (95%), and a smaller subset of T cells express a TCR with gamma (γ) and delta (δ) chains.
The TCR, through its CDRs, endows the T cell with the ability to recognize and respond to foreign or “non self” material. Antigen presenting cells (APCs) digest pathogens and display their fragments on major histocompatibility complex (MHC) molecules. This MHC/antigen complex binds to the TCR while other co-stimulatory molecules (e.g. CD28) are activated leading to T cell activation, proliferation, differentiation, apoptosis, or cytokine release (Samelson 2011). MHC/antigen complexes are, however, not the only molecules capable of interaction with TCRs. Non-peptide antigens such as lipids can interact with TCRs via some of the five isoforms of CD1 (a-e) (Mori and De Libero 2012), and several studies describe TCRs binding to metabolic intermediates bound to the MHC like molecule MR1 (Reantragoon et al. 2012).
TCRs have been described as the most intricate receptor structures of the mammalian immune system (Clambey et al. 2014). In the following sections, we describe how these unique structures are developed, the mechanisms used to drive immune responses via TCR signaling, the effects of their dysfunction and their exploitation as therapeutics.

Fig. 1. Diagram of TCR engagement with the peptide antigen MHC complex using the CD4 T cell as an example.
TCR development and diversity
The TCR is a disulfide-linked membrane bound heterodimeric protein normally consisting of the highly variable α and β chains expressed as part of a complex with the invariant CD3 chain molecules. T cells expressing these two chains are referred to as α:β (or αβ) T cells, though a minority of T cells express an alternate receptor, formed by variable γ and σ chains, referred as γσ T cells. TCR development occurs through a lymphocyte specific process of gene recombination, which assembles a final sequence from a large number of potential segments. This genetic recombination of TCR gene segments in somatic T cells occurs during the early stages of development in the thymus. The TCRα gene locus contains variable (V) and joining (J) gene segments (Vβ and Jβ), whereas the TCRβ locus contains a D gene segment in addition to Vα and Jα segments. Accordingly, the α chain is generated from VJ recombination and the β chain is involved in VDJ recombination. This is similar for the development of γδ TCRs, in which the TCRγ chain is involved in VJ recombination and the TCRδ gene is generated from VDJ recombination (Figure 2). Recombination is temporally regulated by the access of recombinant activating gene (RAG) 1 and RAG2 to recognizable sequences. TCR recombination occurs at two stages during the process of T cell development. First, the β chain gene undergoes Dβ – Jβ rearrangement before Vβ – DJβ recombination in the double negative cells of the thymus. Rearrangement of the α chain gene takes place in double positive thymocytes. RAG1/2 bind to and introduce double strand breaks at recombination signal sequences (RSS), which flank all TCR gene segments. DNA repair machinery completes the recombination reaction. RAG1/2 are expressed by all lymphoid progenitors and immature T and B cells.
The TCR α chain gene locus consists of 46 variable segments, 8 joining segments and the constant region. The TCR β chain gene locus consists of 48 variable segments followed by two diversity segments, 12 joining segments and two constant regions. The D and J segments are located within a relatively short 50 kb region while the variable genes are spread over a large region of 1.5 mega bases (TCRα) or 0.67 megabases (TCRβ). Therefore, the process of VDJ recombination requires contraction of the entire locus by looping. In precursor cells, the gene loci are localized at the periphery of the nucleus where gene transcription is less active. However the proximity allows for D-J recombination at this stage. As differentiation progresses the gene locus moves to a more central nuclear position which likely promotes chromatin opening. As the cells commit to a particular lineage, the variable (V) gene loci undergo a contraction that places usually distant V genes next to the already arranged D-J segments. The contraction produces a looping that allows the different V genes access to the already combined DJ segment with similar frequency.
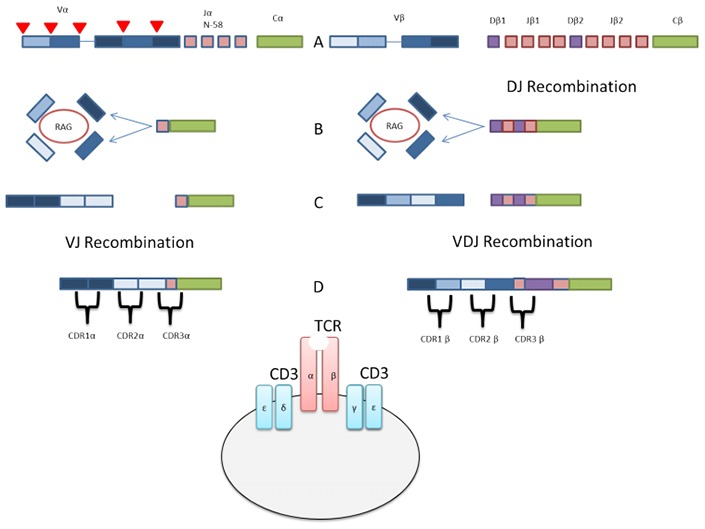
Fig. 2. Simplified overview of VJ & V(D)J recombination of alpha and beta chains of TCRs
A: TCR gene locus showing the (V) variable, (J) joining and (C) constant regions. Red triangles denote the RSS at the edge of each gene segment. DJ recombination of the beta chain occurs.
B: Following DJ recombination in the precursor cells. The locus has moved to a more central location. The V gene segments undergo the looping allowing each at chance at combination with the DJ segments
C: Combination of the selected V regions with DJ segment
D: Mature TCR transcripts showing the approximate location of complementary determining regions (CDRs)
The ultimate goal of the recombination process is diversity among the CDRs. CDRs are the antigen binding sections of TCRs and a diverse recognition capability leads to efficient protection against pathogens and the generation of optimal immune responses. VDJ recombination, along with the addition or deletion of nucleotides at the junctions between gene segments helps to generate an astounding amount of TCR diversity. The diversity generated by VDJ recombination is estimated to exceed 1015 TCRs (Nikolich-Zugich J 2004). CDRs can be re-edited in the periphery, in a process called TCR revision. This results in a change in the antigen specificity of the TCR, and is initiated by reactivation of recombinases that initiate gene recombination (Scott Hale and Fink 2010). TCR revision is a carefully regulated process that induces tolerance to self-antigens, while still inducing protection against invading pathogens.
In addition to the diversity of TCRs, their cross-reactivity also contributes to host defense. Cross-reactivity of TCRs means giving a single T cell host the ability to react to multiple peptide ligands. This is achieved by a degree of flexibility in TCR-MHC/peptide binding. This means fewer T cells are required to detect foreign material. Additionally antigen may be detected by several T cells leading to a polyclonal response, which will make pathogen escape more difficult. This wide ranging cross-reactivity is not without its problems as the concept of T cells being activated by antigen and then cross-reacting to self-ligands is possible. This is known as molecular mimicry and it represents one theory of the root cause of auto-immunity.
TCR activation and signaling
The function of T cells is controlled by TCR activation and signaling. Stimulation of T cell function is initiated upon interaction of the TCR with short peptides presented by MHC class I or II molecules (MHC 1 for CD8 T cells and MHC II for CD4 T cells) (Figure 1). However, the TCR heterodimer by itself is incapable of activating downstream pathways to initiate T cell activation. Initiation of TCR signaling requires co-receptors such as CD4 for helper T cells and CD8 for cytotoxic T cells. These co-receptors act as cellular adhesion molecules that bind their respective MHC molecules and stabilize the interaction of T cells and antigen presenting cells.
The TCR is also located in close proximity to a complex of signaling molecules, which help to mediate T cell activation. These include the CD3 family of proteins (CD3δ, CD3ε, and CD3γ) as well as a TCR zeta (ζ) chain (Figure 1) (Wucherpfennig et al. 2010). Once the TCR is properly engaged with the peptide-MHC complex, conformational changes in the associated CD3 chains are induced, which leads to their phosphorylation and association with downstream proteins (Huse 2009, Alarcòn et al. 2003). The TCR ζ chain is also phosphorylated upon TCR engagement. These molecules are phosphorylated via their c-terminal immunoreceptor tyrosine-based activation motifs (ITAMs) by the Src kinases leukocyte-specific tyrosine kinase (LCK) and Fyn.
Phosphorylated CD3 ITAMs then recruit and activate the Syk family kinase zeta-activated protein 70 kDa (ZAP70) via Src-homology-2 (SH2)-domain interactions (Lin and Weiss 2001). ZAP70 then phosphorylates a membrane associated scaffolding protein called linker for activation of T cells (LAT) (Samelson 2002). LAT in turn recruits a second molecular scaffold, SH2-domain containing leukocyte protein of 76 kDa (Slp-76). Slp-76 is then phosphorylated by ZAP70 and the resulting LAT-Slp-76 complex acts as a scaffold for the recruitment of signaling effector molecules (Koretzky et al. 2006). Interleukin-2 inducible tyrosine kinase (ITK) then interacts with the LAT-Slp-76 complex and becomes activated by autophosphorylation. This promotes phosphorylation of the effector molecule phospholipase C gamma 1 (PLC-γ1). PLC-γ1 transduces TCR signals by cleaving phosphatidylinositol triphosphate (PIP2) in the plasma membrane to generate the second messengers diacylglycerol (DAG) and inositol trisphosphate (IP3).
DAG, a membrane associated lipid, activates a number of downstream proteins, including various isoforms of protein kinase C (PKC) and RAS guanyl nucleotide-releasing protein (RasGRP). Upon activation by DAG, PKC-theta participates in activating the NF-κB pathway (Lin and Weiss 2001), whereas RasGRP is a crucial activator of MAPK signaling pathways (Huse 2009). IP3, on the other hand, stimulates the efflux of Ca2+ from the endoplasmic reticulum to the cytoplasm (Katan 1996). Elevated Ca2+ levels induce activation of the protein phosphatase, calcineurin, which then dephosphorylates the T cell transcription factor nuclear factors of activated T cells (NFAT). Dephosphorylated NFAT then migrates to the nucleus to join other transcription factors in inducing the transcription of specific genes.
In general, the signaling events described ultimately lead to cytoskeletal changes and transcription of relevant T cell genes. However, signaling solely through the TCR is not sufficient for T cell activation, and can result in induction of an anergic state in which T cells fail to respond to antigen stimulation and cannot be restimulated. Therefore, a central tenet of T cell activation is the requirement of co-stimulation through ligation of the TCR with other cell surface receptors which provide additional signals to promote productive T cell activation. Several co-stimulatory molecules have been identified, including CD28, CD2, CD5, CD30, 4-1BB, OX40, LFA-1 and inducible costimulatory (ICOS). CD28-deficient mice demonstrate dampened immune responses to a number of infectious agents (Acuto and Michel 2003). However, not all immune responses are severely impacted by loss of CD28, indicating the dual role of other co-stimulatory molecules in T cell activation (Bachmaier et al. 1996).
TCR signaling is still an active area of research and future studies will undoubtedly reveal new interactions between current proteins involved in TCR signaling as well as identify new proteins and interactions. A summary of the major aspects of TCR signaling is shown in Figure 3.
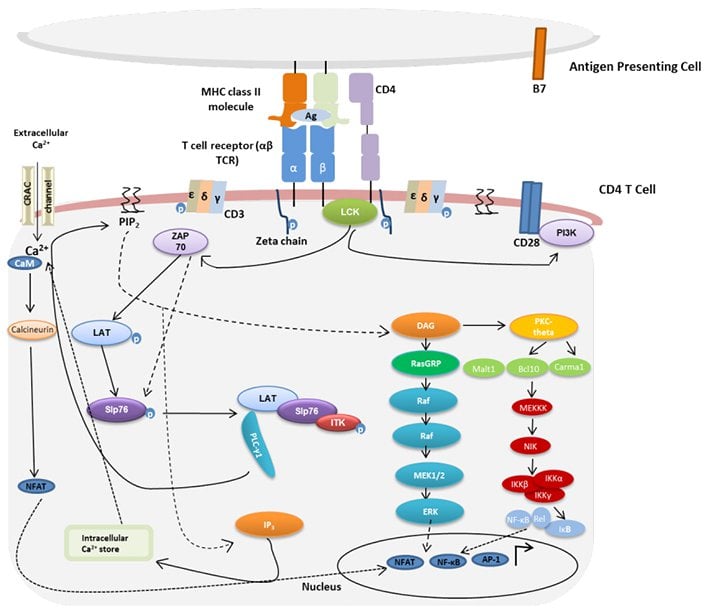
Fig. 3. Overview of TCR signaling. Adapted from Samelson 2011. TCR signaling is activated upon interaction of the TCR with cognate peptide antigen bound to a major histocompatibility complex (MHC) molecule, and co-stimulation by co-receptor molecules such as CD28. AP-1, activator protein 1; Bcl10, B cell lymphoma 10; Ca2+, calcium; CaM, calmodulin; Carma1, caspase recruitment domain membrane-associated guanylate kinase protein 1; CRAC, calcium release-activated Ca2+; DAG, diacylglycerol; IP3, inositol trisphosphate; IKK, I kappa B kinase; ITK, interleukin-2 inducible tyrosine kinase; LAT, linker activation of T cells; LCK, leukocyte-specific tyrosine kinase; Malt 1, mucosa-associated lymphoid tissue protein 1; NFAT, nuclear factors of activated T cells; NF-κB, nuclear factor kappa B; NIK, NF-kappa-B-inducing kinase, PI3K, phosphatidylinositol-3 kinase; PIP2, phosphatidylinositol bisphosphate; PKC-theta, protein kinase C-theta; PLC-γ1, phospholipase C gamma 1; RasGRP, RAS guanyl nucleotide releasing protein; Slp76, SH2-domain containing leukocyte protein of 76 kDa; ZAP70, zeta-activated protein 70 kDa.
Clinical manifestations of aberrant TCR signaling
Genetic defects in critical TCR signaling proteins have been shown to result in distinct clinical and immunological phenotypes. Mutations in ZAP70, LCK and ITK genes have been identified in patients demonstrating various immune pathologies (Notarangelo 2014). Below we describe the reported studies of the clinical manifestations of these defects in humans.
Effect of defects in ZAP70 function
It was in the early 1990s that scientists first discovered that mutations in the ZAP70 gene were directly associated with a rare primary immunodeficiency called CD8 deficiency (Elder et al. 1994, Chan et al. 1994, Arpaia et al. 1994). CD8 deficiency was first characterized as a lack of circulating CD8+ T cells, with the presence of non-functional CD4+ T cells. Studies investigating the mechanism of immune dysregulation associated with ZAP70 defects in humans have shown that CD4+ cells from ZAP-70 deficient patients have an altered T cell repertoire (Roifman et al. 2010). These cells express a phenotype consistent with autoimmunity, which includes reduced levels of CTLA4, IL-10 and TGFβ gene transcripts and decreased sensitivity to FAS-mediated apoptosis (Roifman et al. 2010).
A study by Katamura et al. (1999) described a patient with CD8 deficiency, with a unique subset of CD4 T cells. The patient presented with infiltrative cutaneous erythematous lesions and accumulation of activated CD4+ CD25+ CD45RO+ DR+ T cells and eosinophils in the perivascular area of the skin. Another more recent study showed that ZAP70 missense mutations resulted in a clinical phenotype characterized by immune dysregulation, with wheezing, generalized erythroderma, lymphadenopathy, eosinophilia and elevated serum IgE levels (Turul et al. 2009).
Effects of LCK deficiency
The first report of molecularly-confirmed LCK deficiency in humans was published in 2012. Hauck et al. (2012) described the case of a child who presented with recurrent respiratory infections, nodular skin lesions, arthritis, vasculitis and autoimmune thrombocytopenia (Notarangelo 2014). In addition to these clinical manifestations, the researchers also noted CD4+ T cell lymphopenia, including reduced T-regulatory (Treg) cells, and an increase in exhausted effector memory CD8+ T cells. Genetic analyses identified a missense mutation in the LCK gene, resulting in a defective protein lacking kinase activity. Associated with this defect in LCK function was impaired intracellular signaling in response to TCR/CD3 stimulation. It is likely that the abnormal TCR signaling as well as the low number of Treg cells may have contributed to the induction and perpetuation of the clinical manifestations of immune dysregulation observed.
Effects of ITK deficiency
ITK is critical to TCR signaling and subsequently T cell activation as it promotes PLC-γ1 activation and Ca2+ influx in response to TCR stimulation. ITK is also necessary for maintaining peripheral immune homeostasis. Accordingly, ITK deficiency or impaired function results in severe immune pathology in humans. In recent years, several patients with autosomal recessive ITK deficiency have been reported (Notarangelo 2014). Their clinical phenotype includes infections, primarily due to herpesviruses, autoimmune cytopenias, lymphadenopathy, hepatosplenomegaly and lymphoproliferative disease, particularly of the lungs (Notarangelo 2014). Molecular studies have also demonstrated naïve CD4+ T cell lymphopenia, increased proportion of activated T cells, defective proliferation to CD3 stimulation, deficiency of NKT cells and progressive hypogammaglobulinemia in patients with ITK deficiency (Notarangelo 2014).
In addition to the described consequences of mutated or non-functional ZAP70, LCK and ITK, other examples of altered T cell signaling associated with defects in critical proteins have been described in mice (Notarangelo 2014). Identification of these conditions could lead to the development of novel therapeutics to treat autoimmune diseases.
T cell mediated therapies
T cell immunotherapies are beginning to emerge as potential breakthrough in cancer therapeutics with initial clinical trials reporting high levels of remission (Gardener et al. 2016, Wang et al. 2016). There are several types of existing T cell based immunotherapies.
Adoptive cell therapy (ACT), a technique pioneered by Philip Greenberg, MD, and a team at the Hutchinson Center in the early 1990s is perhaps the most basic and the foundation of other techniques. In this treatment, a patient’s white blood cells are collected in a process called leukapheresis. Isolation of the specific T cells with the most cancer-fighting potency is followed by in vitro culture of the cells. This hugely expanded T cell population is then reinfused into the patient to work against the cancer.
A further twist on the adoptive cell therapy is the Chimeric Antigen Receptor (CARs) based therapy. CARs are engineered receptors, which graft an arbitrary specificity onto an immune effector cell. Typically, these receptors are used to graft the specificity of a monoclonal antibody onto a T cell, with transfer of their coding sequence facilitated by retroviral vectors. The receptors are called chimeric because they are composed of parts from different sources. First generation CARs typically had the intracellular domain from the CD3 ζ- chain, which is the primary transmitter of signals from endogenous TCRs. Second generation CARs add intracellular signaling domains from various costimulatory protein receptors (e.g., CD28, 41BB, ICOS) to the cytoplasmic tail of the CAR to provide additional signals to the T cell. Cell based therapies are not without challenges; particularly their ability to induce large scale damaging and non-specific immune responses.
Soluble T cell receptor therapy represents a potential alternative to cell based immunotherapies. Bispecific T cell engaging TCR can recognize tumor peptides on the cell surface. This engagement can be used to target therapeutics to this cell or engage further T cell responses.
References
- Acuto O and Michel F (2003). CD28-mediated co-stimulation: a quantitative support for TCR signaling. Nat Rev Immunol 3, 939-951.
- Alarcón B et al. (2003). Initiation of TCR signaling: regulation within CD3 dimers. Immunol Rev 191, 38-46.
- Arpaia E et al. (1994). Defective T cell receptor signaling and CD8+ thymic selection in humans lacking zap-70 kinase. Cell 76, 947-958.
- Bachmaier K et al. (1996). Induction of autoimmunity in the absence of CD28 costimulation. J Immunol 157, 1752-1757.
- Chan AC et al. (1994). ZAP-70 deficiency in an autosomal recessive form of severe combined immunodeficiency. Science 264, 1599-1601.
- Clambey ET et al. (2014). Molecules in medicine mini-review: the αβ T cell receptor. J Mol Med (Berl) 92, 735-741.
- Elder ME et al. (1994). Human severe combined immunodeficiency due to a defect in ZAP-70, a T cell tyrosine kinase. Science 264, 1596-1599.
- Gardner R et al. (2016). Acquisition of a CD19-negative myeloid phenotype allows immune escape of MLL-rearranged B-ALL from CD19 CAR-T-cell therapy. Blood 127, 2406-2410.
- Hauck F et al. (2012). Primary T-cell immunodeficiency with immunodysregulation caused by autosomal recessive LCK deficiency. J Allergy Clin Immunol 130, 1144-1152.
- Huse M (2009). The T-cell-receptor signaling network. J Cell Sci 122, 1269-1273.
- Katamura K et al. (1999). Existence of activated and memory CD4+ T cells in peripheral blood and their skin infiltration in CD8 deficiency. Clin Exp Immunol 115, 124-130.
- Katan M (1996). The control of inositol lipid hydrolysis. Cancer Surv 27, 199-211.
- Koretzky GA et al. (2006). SLP76 and SLP65: complex regulation of signalling in lymphocytes and beyond. Nat Rev Immunol 6, 67-78.
- Lin J and Weiss A (2001). T cell receptor signaling. J Cell Sci 114(Pt 2), 243-244.
- Nikolich-Zugich J et al. (2004). The many important facets of T-cell repertoire diversity. Nat Rev Immunol 4, 123-132.
- Notarangelo LD (2014). Immunodeficiency and immune dysregulation associated with proximal defects of T cell receptor signaling. Curr Opin Immunol 31, 97-101.
- Mori L and De Libero G (2012). T cells specific for lipid antigens. Immunol Res 53, 191-199.
- Reantragoon R et al. (2012). Structural insight into MRI-mediated recognition of the mucosal associated invariant T cell receptor. J Exp Med 209, 761- 774.
- Roifman CM et al. (2010). Characterization of zeta-associated protein, 70 kd (ZAP70)-deficient human lymphocytes. J Allergy Clin Immunol 126, 1226- 1233.
- Samelson LE (2002). Signal transduction mediated by the T cell antigen receptor: the role of adaptor proteins. Annu Rev Immunol 20, 371-394.
- Samelson LE (2011). Immunoreceptor signaling. Cold Spring Harb Perspect Biol 3, a011510.
- Scott Hale J and Fink PJ (2010). T-cell receptor revision: friend or foe? Immunology 129, 467-473.
- Smith-Garvin JE et al. (2009). T cell activation. Annu Rev Immunol 27, 591-619.
- Turul T et al. (2009). Clinical heterogeneity can hamper the diagnosis of patients with ZAP70 deficiency. Eur J Pediatr 168, 87-93.
- Wang X et al. (2016). Phase I studies of central-memory-derived CD19 CAR T cell therapy following autologous HSCT in patients with B-cell NHL. Blood 127, 2980-2990.
- Wucherpfennig KW et al. (2010). Structural biology of the T-cell receptor: Insights into receptor assembly, ligand recognition, and initiation of signaling. Cold Spring Harb Perspect Biol 2, a005140.








