Transcription Factors

- On This Page
- Antibodies for the analysis of transcription factors
- Groups / Classes
- Our transcription factor antibody range
Antibodies for the Analysis of Transcription Factors
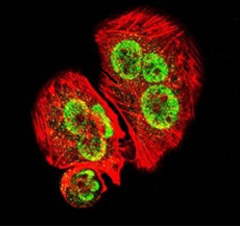
U251 cells were probed with GATA1 polyclonal antibody (AHP2401) (green) and counterstained with phalloidin (F-actin, red)
Transcription factors (TFs) along with RNA splicing, siRNA and histone modifications regulate the translatome. TFs are identifiable through their expression of a specific DNA binding domain (DBD). TFs, via their DBD, recognize and bind to a particular sequence of DNA. TFs work by promoting or blocking the recruitment of RNA polymerase. TFs commonly possess several functional domains which along with DNA binding facilitate interactions with other TFs, coactivators, chromatin, and small non-coding RNA.
TFs are often the apex or final readout of cell signaling pathways. TF activity is regulated by several means including phosphorylation, ligand binding and cellular localization. Phosphorylation of a TF can induce conformational changes in the protein allowing it to bind other TFs or co-factors which leads to transcriptional changes. Binding of cell permeable ligands (e.g. estrogen) to nuclear receptor TFs (e.g. estrogen receptor) can also induce the transcriptional effect of the TFs. Upon phosphorylation and other post translational modifications TFs can alter their cellular localization thereby enabling or preventing their transcriptional effects.
Groups / Classes
Transcription factors are grouped according the DNA binding domains they express. The major groups are as follows:
Classes of Transcription Factors |
Examples |
|---|---|
|
Superclass: basic domains |
bZIP, bHLH, bHLH-ZIP, NF-1, RF-X, bHSH |
|
Superclass: zinc-coordinating DNA-binding domains |
Cys4 zinc finger of nuclear receptor type, diverseCys4 zinc fingers |
|
Superclass: helix-turn-helix |
Paired box, fork head/winged helix, heat shock factors |
|
Superclass: beta-scaffold factors with minor groove contacts |
RHR, STAT, p53, TATA binding proteins, grainyhead, runt |
|
Superclass: other transcription factors |
HMGI(Y)(HMGA1), E1A-like factors, AP2/EREBP-related factors |
The vital role played by TFs is illustrated by their implication in several diseases including cancer. The most common oncogenes and tumor suppressors are TFs including c-myc, p53, NFkB, and STATs.
Our Transcription Factor Antibody Range
| Description | Target | Format | Clone | Applications | Citations | Code |
|---|
- Growth factors
- EGF R resources
- Kinases
- mTOR signaling pathway
- Translation factors
- Phosphatases
- Phosphatases in cancer
- Phospho-Specific precisionAb antibodies
- Phospho-specific antibodies
- Sumoylation mediated signaling
- Ubiquitination mediated signaling
- Tyrosine phosphorylation of EGF R
- Signal transduction antibodies


