Popular topics

-
References
Couzin-Frankel J (2013). Breakthrough of the year 2013. Cancer immunotherapy. Science 342, 1432-1343.
Dong H et al. (2002). Tumor-associated B7-H1 promotes T-cell apoptosis: a potential mechanism of immune evasion. Nat Med 8, 793-800.
Drake JR et al. (2010). Phase I study of single-agent anti-programmed death-1 (MDX-1106) in refractory solid tumors: safety, clinical activity, pharmacodynamics, and immunologic correlates. J Clin Oncol 28, 3167-3175.
Ishida Y et al. (1992). Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death, EMBO J 1, 3887-3895.
Kim ES et al. (2015). Immune checkpoint modulators: An emerging antiglioma armamentarium. J Immunol Res article ID: 4683607.
LaFleur MW et al. (2018). Inhibitors of the PD-1 Pathway in Tumor Therapy. J Immunol 200, 375-383.
McCathy EF (2006). The Toxins of William B. Coley and the Treatment of Bone and Soft-Tissue Sarcomas. IOWA Orthop J 26, 154-158.
Rotte A et al .(2017). Mechanistic overview of immune checkpoints to support the rational design of their combinations in cancer immunotherapy. Annals of Oncology 1, 71-83.
Sharma P and Allison JP (2015). Immune checkpoint targeting in cancer therapy: toward combination strategies with curative potential. Cell 161, 205-214.
Checkmate: Beating Cancer through the PD-1 Pathway
In order to survive, cancer cells need to evade the host immune system. Rather than targeting the tumor itself, immunotherapy aims to enhance the host immune system to more effectively detect and kill cancer cells. Named ‘Breakthrough of the Year’ by Science in 2013 (Couzin-Frankel 2013) the field has since rapidly grown, with accelerated clinical trials demonstrating impressive results. Drugs that target immune checkpoints to enhance the potency of the immune system continue to be a hot topic, despite only a limited number of patients being suited to this therapy.
This blog discusses the PD-1 (programmed cell death-protein 1) pathway and two licensed drugs that target it: pembrolizumab (Keytruda) and nivolumab (Opdivo).
Improving the Potency of the Immune System
William Coley is widely acknowledged as the father of the immunotherapy field. His research showed it was possible to harness a patient’s own immune system to make it more effective at fighting cancer. In the 1890’s he experimented on patients with sarcomas, injecting them with a mixture of Streptococcal bacteria, resulting in their tumors shrinking (McCathy 2006). Over 40 years he injected more than 1000 cancer patients with ‘Coley’s Toxins’ (bacteria or bacterial products) producing good results.
However, the field of immunotherapy wasn’t properly born until the molecular mechanisms governing immune responses were better understood and it was conceivable to try and design a therapy that capitalizes on this knowledge.
Keeping the Immune System in Check
Inappropriate activation of the immune system can result in a variety of autoimmune disorders. Immune checkpoints are crucial for self-tolerance and act to prevent inappropriate activation of the immune system or to halt or dampen down an immune response.
Cancerous cells sometimes exploit these checkpoints to escape destruction and therefore targeting them with a drug has proved an effective therapy for certain cancer types. The most commonly targeted checkpoints are CTLA-4, TIGIT, LAG-3 and PD-1 (Rotte et al. 2017).
The PD-1/PD-L1 Pathway
PD-1 was discovered as a molecule expressed on dying T cells by Tsuku Hokjo’s group in the 1990’s (Ishida et al, 1992). This pathway is important for dampening down the T cell response, and inappropriate activation of this pathway by cancer cells provides an opportunity of immune escape. Indeed, in 2002, Lieping Chen’s group demonstrated that mouse tumors with forced expression of PD-L1 were able to evade the immune system and that the expression of PD-L1 is upregulated in many cancers (Dong et al. 2002). Since tumors expressing high levels of PD-1 are associated with poor outcomes, targeting this pathway can potentially have a dramatic impact on survival.
PD-1 is expressed on activated T cells and its ligands (PD-L1, and PD-L2) are presented by self cells. In the absence of PD-L1/L2 binding, signaling through ZAP-70 and LCK leads to increased T cell proliferation, activation and effector function. Ligand binding results in phosphorylation of PD-1 and the recruitment of protein tyrosine phosphatases (in particular SHP-2). These then dephosphorylate and inactivate downstream signaling molecules such as ZAP-70 and LCK (Figure 1).
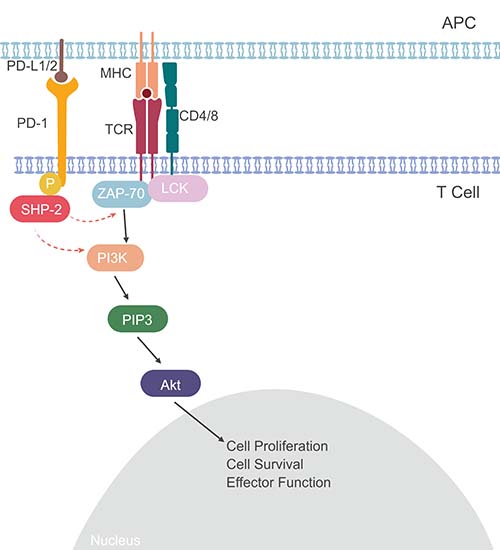
Fig. 1. The PD-1 Pathway. Binding of PD-L1/2 to the PD-1 receptor leads to phosphorylation of SHP-2 and inhibition of signaling through ZAP-70/LCK, and a reduction of T cell activation and effector functions. Red dotted arrows show inhibition, black arrows show activation.
Checkpoint Inhibitors
Monoclonal antibodies have been designed to block the PD-1 pathway (Figure 2) and have shown impressive results in the clinic. The first trial of a PD-1 antibody (MDX-1106, later called nivolumab) saw encouraging results - for some volunteers with end stage disease their tumors shrunk, significantly extending their expected survival (Drake et al. 2010).
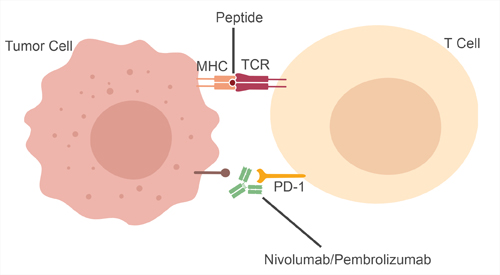
Fig. 2. Mechanism of action of nivolumab and pembrolizumab.
Results like this for both nivolumab and another anti-PD-1 antibody, pembrolizumab resulted in both drugs being licensed by the FDA for refractory melanoma in 2014. (LaFleur et al. 2018).
Further clinical trials have demonstrated the broad therapy potential of this pathway and resulted in nivolumab and pembrolizumab being licensed for treatment of additional cancer types including cervical cancer, refractory primary mediastinal large B-cell lymphoma and lung cancer. More recently, pembrolizumab became the first drug to be granted approval across cancers based on a predictive genetic biomarker, which for therapy approval, is for tumors demonstrating microsatellite instability (LaFleur et al. 2018).
The success of monoclonal antibodies in blocking these checkpoints highlights the critical role they play in regulating T cell responses. However, not every patient is suitable for this treatment and research to identify the best predictive biomarkers is still ongoing.
The Role of Immune Checkpoints in Immunity and Cancer
Our popular mini-review discusses the key role immune checkpoints play in regulating immune activation. Discover more about the major immune checkpoints and how understanding these pathways is leading to advances in the clinic.
Read it NowReferences
Couzin-Frankel J (2013). Breakthrough of the year 2013. Cancer immunotherapy. Science 342, 1432-1343.
Dong H et al. (2002). Tumor-associated B7-H1 promotes T-cell apoptosis: a potential mechanism of immune evasion. Nat Med 8, 793-800.
Drake JR et al. (2010). Phase I study of single-agent anti-programmed death-1 (MDX-1106) in refractory solid tumors: safety, clinical activity, pharmacodynamics, and immunologic correlates. J Clin Oncol 28, 3167-3175.
Ishida Y et al. (1992). Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death, EMBO J 1, 3887-3895.
Kim ES et al. (2015). Immune checkpoint modulators: An emerging antiglioma armamentarium. J Immunol Res article ID: 4683607.
LaFleur MW et al. (2018). Inhibitors of the PD-1 Pathway in Tumor Therapy. J Immunol 200, 375-383.
McCathy EF (2006). The Toxins of William B. Coley and the Treatment of Bone and Soft-Tissue Sarcomas. IOWA Orthop J 26, 154-158.
Rotte A et al .(2017). Mechanistic overview of immune checkpoints to support the rational design of their combinations in cancer immunotherapy. Annals of Oncology 1, 71-83.
Sharma P and Allison JP (2015). Immune checkpoint targeting in cancer therapy: toward combination strategies with curative potential. Cell 161, 205-214.
You may also be interested in...

View more cancer or science-news blogs