p53 Antibody: An Introductory Guide
Structure, Isoforms, Application Information, Phosphorylation and Apoptosis
The p53 tumor suppressor/transcription factor regulates a wide variety of cellular functions, including apoptosis, cell cycle, DNA repair and metabolism. Since its discovery in 1979, p53 has been the subject of intense scrutiny. However, such is the complex behavior of this protein, it continues to challenge researchers despite nearly four decades of research.
This introductory guide aims to provide useful information for researchers beginning their p53 antibody based experiments.
View our Full Range of p53 Antibodiesp53 Structure
The gene that encodes p53 is found on chromosome 17. The fullest length isoform produces a protein of 393 amino acids (53 kDa). Figure 1 is a simple schematic of the p53 structure and highlights the epitope for several of the most popular p53 antibodies.
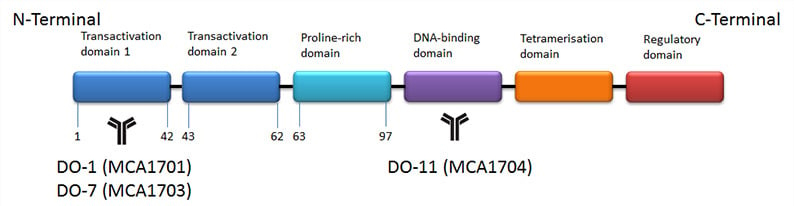
Fig. 1.
Structure
of p53.
 indicates the location of the epitope of the antibodies listed below.
indicates the location of the epitope of the antibodies listed below.
p53 Isoforms
There are 12 isoforms of p53 created by alternative translation sites, alternative splicing and through the use of different promoters. The structures of the individual isoforms are shown in Figure 2. p53 isoforms are expressed in a tissue dependent manner and full length p53 is never expressed alone.
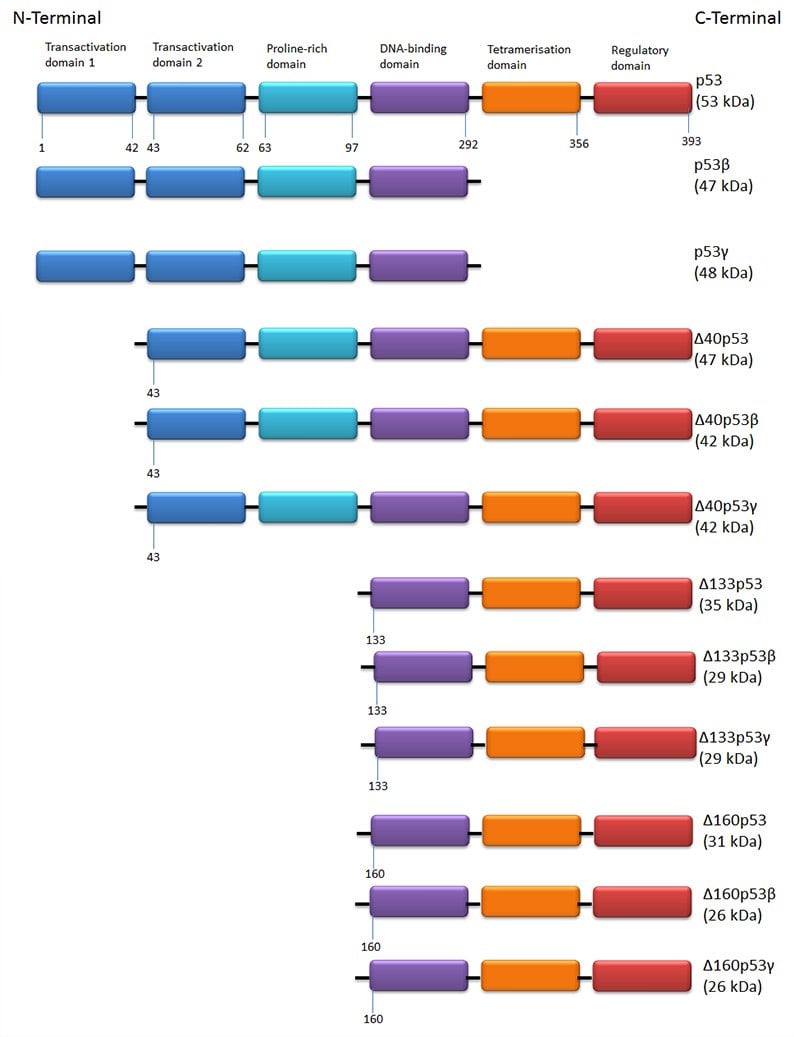
Fig. 2. The 12 isoforms of p53. Full length p53, p53β and p53γ differ in the expression of C terminal tetramerization domains. The ?40 group have the same C-terminal variation but have lost the first 39 amino acids from the transactivation domain. Similarly the ?133 and ?160 groups lack significant portions of the N-terminal end of the protein.
Depending on the location of the epitope, antibodies may detect different isoforms of p53. Table 1 below lists the isoforms detected by three of the best characterized antibodies.
Table 1. p53 Antibodies.
Clone |
Isoforms Detected |
Applications |
Formats |
Catalog # |
|---|---|---|---|---|
|
DO-1 |
p53, p53β, p53γ |
FC, IHC-F, E, IP, IHC-P, WB |
Pur., FITC, PE, B. |
|
|
DO-7 |
p53, p53β, p53γ |
IHC-F, FC, IP, IHC-P, WB |
Pur. |
|
|
DO-11 |
All isoforms |
IHC-F, IP, IHC-P, WB |
Pur. |
B., biotin; FC, Flow Cytometry; E, ELISA; IHC-F, immunohistochemistry frozen; IHC-P, immunohistochemistry paraffin ; IP, immunoprecipitation; PE, phycoerythrin; Pur., purified; WB, western blot.
Application Specific Approaches for Studying p53
Under normal conditions p53 has a short half-life and is subject to degradation by ubiquitin dependent systems. p53 is stabilized and activated in response to a range of cellular stresses including metabolic stress and DNA damage. This is mediated mainly through uncoupling p53 from its key negative regulators, MDM2 and MDM4, leading to the accumulation of stable active p53. Therefore, when studying p53, attention must be paid to the expression system selected. Often detection of p53 may require exposure of cells to stressful conditions such as UV light.
Western blotting
Due its relevant abundance, p53 is detectable by western blot without the need for specialized protocols. What may vary is the number of isoforms detected by the primary antibody. Table 1 above lists the isoforms detected by three of the most popular clones. Researchers may wish to specifically detect nuclear accumulation of p53 as this is particularly indicative of stress response. For detecting nuclear proteins, Bio-Rad recommends the ReadyPrep™ Protein Extraction Kit (Cytoplasmic/Nuclear).
Flow cytometry
Clones DO-1 (MCA1701) and DO-7 (MCA1703) are validated in flow cytometry. However, due to the intracellular location of p53, analysis by flow cytometry requires cell permeabilization. To assist with p53 flow cytometry experiments, Bio-Rad offers a wide variety of supporting reagents including cell permeabilization reagents.
Immunohistochemistry
DO-1, DO-7 and DO-11 are validated in IHC of both frozen and paraffin sections. Detection of p53 in paraffin sections using DO-1, DO-7 or DO-11 requires antigen retrieval using heat treatment prior to staining of paraffin sections. Sodium citrate buffer pH 6.0 is recommended for this purpose. Colon or breast carcinoma tissue is recommended as a positive control tissue. For more information on IHC protocols, visit the IHC resource page.
Phosphorylation of p53
Cell stressors (e.g. UV light) promote a series of reversible post-translational modifications (PTMs) of p53 including multisite phosphorylation of the transactivation domain (N-terminus). Figure 3 shows the location of the phosphorylation sites of full length p53. Table 2 provides details of the kinases responsible for each phosphorylation as well as antibodies specific to each site where possible.
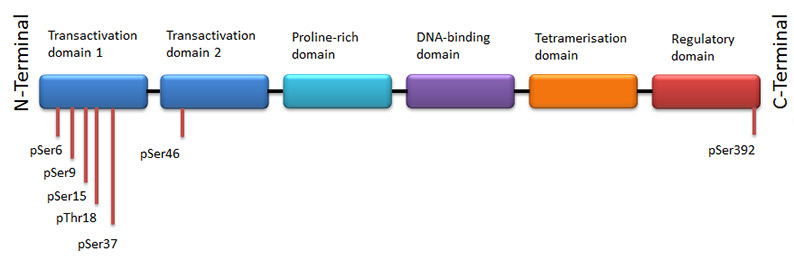
Fig. 3. Phosphorylation sites of p53.
Table 2. Phosphorylation sites of p53.
Phosphorylation Site |
Kinases Responsible for Phosphorylation |
|---|---|
|
pSer6 |
Casein kinase 1-delta |
|
pSer9 |
Casein kinase 1-delta, DNA-dependent protein kinase |
|
pThr18 |
DNA-dependent protein kinase, casein kinase 1 |
|
pSer392 |
Casein kinase 2 |
p53 and Apoptosis
Upon apoptotic stimulation, p53 translocates from the cell nucleus to the mitochondria, where it interacts with B cell lymphoma 2 (Bcl-2) family of proteins to regulate apoptosis. p53 binds to the pro-survival members, liberating the multi-domain pro-apoptotic members that then trigger the mitochondria related stages of apoptosis. Table 3 lists several of the pro-apoptotic and anti-apoptotic Bcl-2 family members. Table 4 lists antibodies against proteins relevant in p53 mediated apoptosis. For more information on apoptosis, refer to the Apoptosis page.
Table 3. Bcl-2 family members.
|
Anti-apoptotic proteins |
Bcl-2, Bcl-xL, Bcl-w, A1 and Mcl-1 The survival activity of the anti-apoptotic proteins may be blocked by the binding of Bcl-2 family of pro-apoptotic proteins. |
|
Pro-apoptotic proteins |
BH3 only subfamily – BAD, BID, BIM, Bik, Blk, Hrk, BNIP3 , and BimL Multi-domain subfamily – BAX BAK and BOK |
Bcl-xL, Bcl-2-like protein 1 long isoform; Bcl-w, BCL2L2; BAD,Bcl2-associated agonist of cell death; BID, BH3 interacting domain death agonist; BIM, BCL2L11; Bik, Bcl-2-interacting killer; BNIP3, BCL2/adenovirus E1B 19 kDa protein-interacting protein 3; BAX, Bcl-2 like protein 4; BAK, Bcl-2 homologous antagonist/killer; BOK, Bcl-2 related ovarian killer.
Table 4. Antibodies against proteins involved in p53 mediated apoptosis.
Target |
Synonyms |
Catalog # |
Further Information |
|---|---|---|---|
|
bcl-2 oncoprotein |
MCA1550 |
||
|
Bcl-2-like protein 1 (long isoform) |
AHP2558
|
pSer62 inhibits anti-apoptotic activity of Bcl-xL |
|
|
Bcl-2 like protein 4 |
AHP2716 |
||
|
p53 upregulated modulator of apoptosis |
AHP727 |
|
For further information on tumor suppressors, visit the tumor suppressor page or view a complete list of tumor suppressor antibodies.




