Poly(ADP-Ribose) Polymerase-1 antibody | A6.4.12




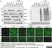


Mouse anti Poly(ADP-Ribose) Polymerase-1
- Product Type
- Monoclonal Antibody
- Clone
- A6.4.12
- Isotype
- IgG1
- Specificity
- Poly(ADP-Ribose) Polymerase-1
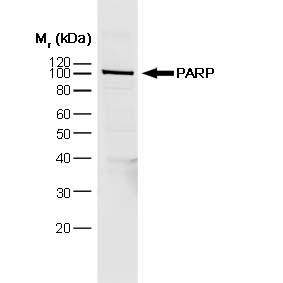
| Mouse anti poly (ADP-ribose) polymerase 1 antibody, clone A6.4.12 recognizes poly (ADP-ribose) polymerase 1 (PARP-1), a ~116 kDa nuclear enzyme, cleaved during apoptosis (Soldani et al. 2002). PARP-1, a caretaker enzyme, is involved in DNA damage repair (Langelier et al. 2013), plays roles in diabetes pathophysiology (Andreone et al. 2012) and tumour proliferation ( As well as protecting cells from genomic instability, PARP-1 is involved in the development of both inflammatory and immune responses, and cell death by apoptosis and necrosis (Erdélyi et al. 2005). Mouse anti poly(ADP-ribose) polymerase 1 antibody, clone A6.4.12, targets PARP-1, an enzyme which represents a promising target for new developments in therapeutic treatment of immune mediated diseases ( |
- Target Species
- Human
- Species Cross-Reactivity
-
Target Species Cross Reactivity Hamster Mouse Drosophila Xenopus Rat - N.B. Antibody reactivity and working conditions may vary between species.
- Product Form
- Purified IgG - liquid
- Preparation
- Purified IgG prepared by affinity chromatography on Protein A from tissue culture supernatant
- Buffer Solution
- Phosphate buffered saline
- Preservative Stabilisers
- 0.09% sodium azide (NaN3)
- Carrier Free
- Yes
- Immunogen
- Human PARP-1
- Approx. Protein Concentrations
- IgG concentration 1.0 mg/ml
- Fusion Partners
- Spleen cells from immunized BALB/c mice were fused with cells of mouse NS0 myeloma cell line.
- Regulatory
- For research purposes only
- Guarantee
- 12 months from date of despatch
Avoid repeated freezing and thawing as this may denature the antibody. Storage in frost-free freezers is not recommended.
| Application Name | Verified | Min Dilution | Max Dilution |
|---|---|---|---|
| ELISA | |||
| Immunofluorescence | |||
| Immunohistology - Frozen | |||
| Immunohistology - Paraffin 1 | |||
| Immunoprecipitation | |||
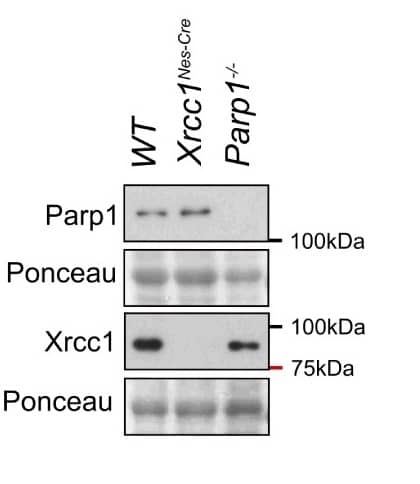
| Western Blotting | 1/1000 | 1/5000 |
- 1 Clone A6.4.12 requires antigen retrieval using heat treatment prior to staining of paraffin sections. Sodium citrate buffer pH 6.0 is recommended for this purpose.
References for Poly(ADP-Ribose) Polymerase-1 antibody
-
Harris, J.L. et al. (2009) Aprataxin, poly-ADP ribose polymerase 1 (PARP-1) and apurinic endonuclease 1 (APE1) function together to protect the genome against oxidative damage.
Hum Mol Genet. 18: 4102-17. -
Freire, R. et al. (2001) Cleavage of the Bloom's syndrome gene product during apoptosis by caspase-3 results in an impaired interaction with topoisomerase IIIalpha.
Nucleic Acids Res. 29 (15): 3172-80. -
Krohn, A.J. et al. (1998) Staurosporine-induced apoptosis of cultured rat hippocampal neurons involves caspase-1-like proteases as upstream initiators and increased production of superoxide as a main downstream effector.
J Neurosci. 18 (20): 8186-97. -
Staples, C.J. et al. (2010) Cross-talk between the p38alpha and JNK MAPK pathways mediated by MAP kinase phosphatase-1 determines cellular sensitivity to UV radiation.
J Biol Chem. 285 (34): 25928-40. -
Alexander, B.M. et al. (2010) DNA repair protein biomarkers associated with time to recurrence in triple-negative breast cancer.
Clin Cancer Res. 16: 5796-804. -
Gueven, N. et al. (2004) Aprataxin, a novel protein that protects against genotoxic stress.
Hum Mol Genet. 13 (10): 1081-93. -
Gueven, N. et al. (2006) Defective p53 response and apoptosis associated with an ataxia-telangiectasia-like phenotype.
Cancer Res. 66: 2907-12. -
Kim, J.W. et al. (2000) Inhibition of homodimerization of poly(ADP-ribose) polymerase by its C-terminal cleavage products produced during apoptosis.
J Biol Chem. 275: 8121-5.
View The Latest Product References
-
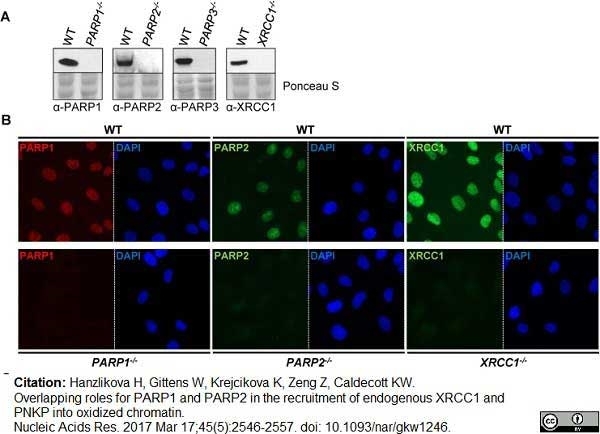
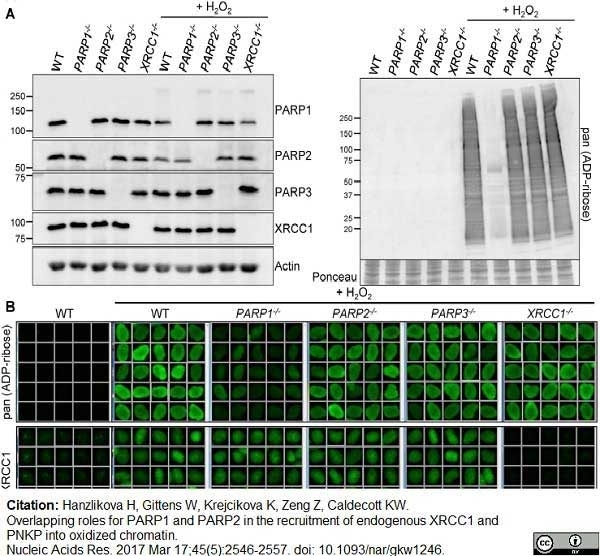
Hanzlikova, H. et al. (2017) Overlapping roles for PARP1 and PARP2 in the recruitment of endogenous XRCC1 and PNKP into oxidized chromatin.
Nucleic Acids Res. 45 (5): 2546-2557. -
Olaussen, K.A. et al. (2013) PARP1 impact on DNA repair of platinum adducts: Preclinical and clinical read-outs.
Lung Cancer. 80: 216-22. -
Fabrice, A.
et al . (2012) PARP and adjuvant cisplatin-based chemotherapy in non-small-cell lung cancer.
US patent: 20120277110 -
Geistrikh, I. et al. (2011) Ca2+-induced PARP-1 activation and ANF expression are coupled events in cardiomyocytes.
Biochem J. 438: 337-47. -
Mirzaa, G.M. et al. (2014) Mutations in CENPE define a novel kinetochore-centromeric mechanism for microcephalic primordial dwarfism.
Hum Genet. 133: 1023-39. -
Milner, R. et al. (2013) Validation of the BRCA1 antibody MS110 and the utility of BRCA1 as a patient selection biomarker in immunohistochemical analysis of breast and ovarian tumours.
Virchows Arch. 462: 269-79. -
Inbar, D. et al. (2012) Erythropoietin-driven signalling and cell migration mediated by polyADP-ribosylation.
Br J Cancer. 107: 1317-26. -
Buchsbaum, S. et al. (2012) FAT10 is a proteasomal degradation signal that is itself regulated by ubiquitination.
Mol Biol Cell. 23: 225-32. -
Mullane, S.A. et al. (2016) Expression Levels of DNA Damage Repair Proteins Are Associated With Overall Survival in Platinum-Treated Advanced Urothelial Carcinoma.
Clin Genitourin Cancer. 14 (4): 352-9. -
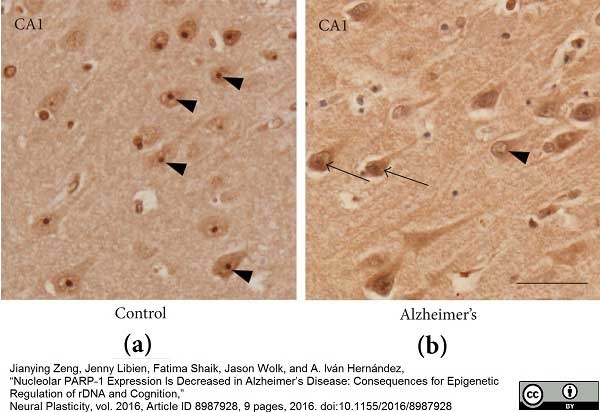
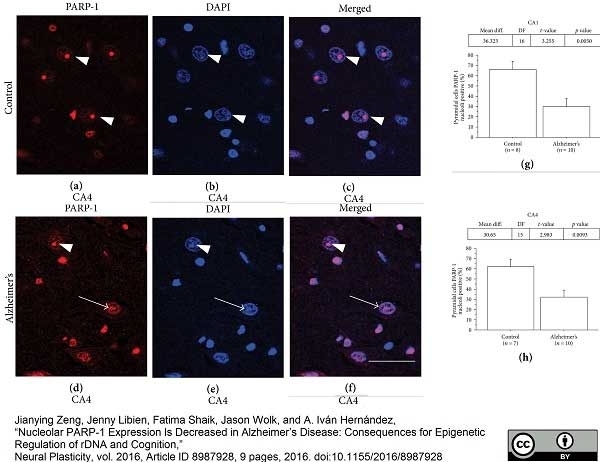
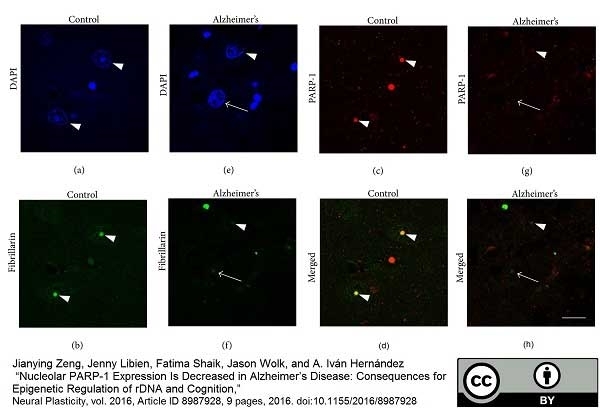
Zeng, J. et al. (2016) Nucleolar PARP-1 Expression Is Decreased in Alzheimer's Disease: Consequences for Epigenetic Regulation of rDNA and Cognition.
Neural Plast. 2016: 8987928. -
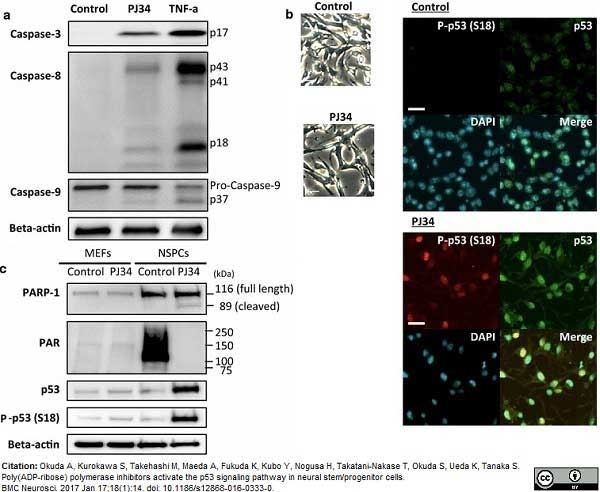
Okuda, A. et al. (2017) Poly(ADP-ribose) polymerase inhibitors activate the p53 signaling pathway in neural stem/progenitor cells.
BMC Neurosci. 18 (1): 14. -
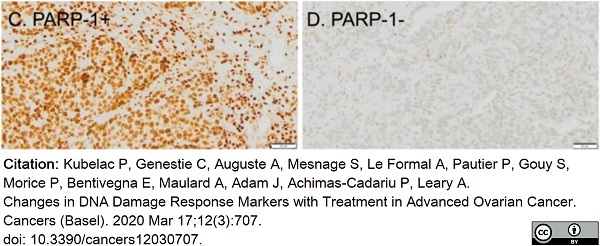
Kubelac, P. et al. (2020) Changes in DNA Damage Response Markers with Treatment in Advanced Ovarian Cancer.
Cancers (Basel). 12(3): 707. -
Komulainen, E. et al. (2021) Parp1 hyperactivity couples DNA breaks to aberrant neuronal calcium signalling and lethal seizures.
EMBO Rep. 22 (5): e51851. -
Rosso, I. et al. (2023) Alternative lengthening of telomeres (ALT) cells viability is dependent on C-rich telomeric RNAs.
Nat Commun. 14 (1): 7086.
Further Reading
-
Pinton, G. et al. (2013) PARP1 inhibition affects pleural mesothelioma cell viability and uncouples AKT/mTOR axis via SIRT1.
J Cell Mol Med. 17: 233-41. -
Rosado, M. et al. (2013) Beyond dna repair,the immunological role of parp-1 and its siblings.
Immunology. 139: 428-37. -
Andreone, T. et al. (2012) Cytokine-mediatedβ-cell damage in PARP-1-deficient islets.
Am J Physiol Endocrinol Metab. 303: E172-9. -
Langelier, M.F. and Pascal, J.M. (2013) PARP-1 mechanism for coupling DNA damage detection to poly(ADP-ribose) synthesis.
Curr Opin Struct Biol. 23: 134-43.
- RRID
- AB_2236751
- UniProt
- P09874
- Entrez Gene
- PARP1
- GO Terms
- GO:0003677 DNA binding
- GO:0003950 NAD+ ADP-ribosyltransferase activity
- GO:0005635 nuclear envelope
- GO:0005667 transcription factor complex
- GO:0005730 nucleolus
- GO:0006366 transcription from RNA polymerase II promoter
- GO:0006471 protein ADP-ribosylation
- GO:0008134 transcription factor binding
- GO:0008270 zinc ion binding
- View More GO Terms
- GO:0042802 identical protein binding
- GO:0032869 cellular response to insulin stimulus
- GO:0045449 regulation of transcription
- GO:0047485 protein N-terminus binding
MCA1522G
If you cannot find the batch/lot you are looking for please contact our technical support team for assistance.
Please Note: All Products are "FOR RESEARCH PURPOSES ONLY"
View all Anti-Human ProductsAlways be the first to know.
When we launch new products and resources to help you achieve more in the lab.
Yes, sign me up