The Intrinsic Mitochondrial Apoptosis Pathway
The Intrinsic Mitochondrial Apoptosis Pathway
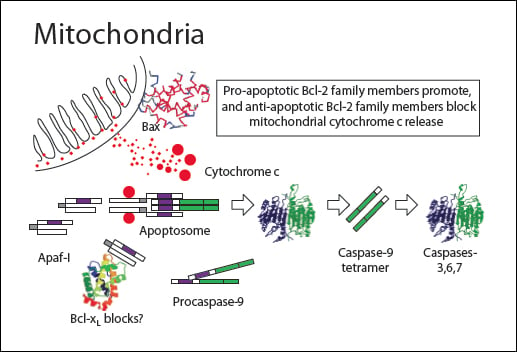 A variety of stress stimuli including growth factor withdrawal, heat shock, and DNA damage activate the apoptosis intrinsic or mitochondrial pathway. Mitochondria are involved in both caspase-dependent and caspase-independent apoptosis pathways. Oxidative damage through the generation of endogenous reactive oxygen species (ROS) by mitochondria can lead to plasma membrane alterations as part of the apoptotic degradation phase.
A variety of stress stimuli including growth factor withdrawal, heat shock, and DNA damage activate the apoptosis intrinsic or mitochondrial pathway. Mitochondria are involved in both caspase-dependent and caspase-independent apoptosis pathways. Oxidative damage through the generation of endogenous reactive oxygen species (ROS) by mitochondria can lead to plasma membrane alterations as part of the apoptotic degradation phase.
View our range of apoptosis antibodies and kits available in different formats and suitable for many applications.
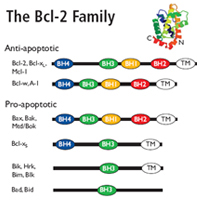
Bcl-2 family
The Bcl-2 family regulates mitochondrial outer membrane permeabilization (MOMP).
Pro-apoptotic Bcl-2 proteins form mitochondrial apoptosis-induced channels (MAC) in the outer mitochondrial membrane.
These mitochondrial membrane pores cause MOMP, releasing cytochrome c and other cytotoxic proteins into the cytosol.
Cytochrome c
Cytochrome c interacts with the cytosolic adapter protein Apaf-1 which recruits initiator procaspase-9. These proteins form a multimeric caspase-activating complex called the apoptosome. Apaf-1 and procaspase-9 bind through a shared homotypic interaction motif called the caspase recruitment domain (CARD). The apoptosome cleaves and activates procaspase-9 into caspase 9 which triggers the caspase cascade by activating the effector caspases 3, caspase 6 and caspase 7. Many proteins are degraded by caspase 3, caspase 6 and caspase 7 causing morphological and biochemical damage with eventual cell death.
Other proteins released from mitochondria into the cytosol after MOMP include:
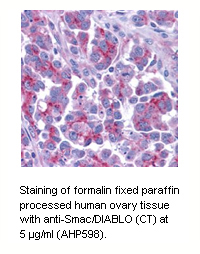
Smac/Diablo and HtrA2/OMI, which are negative regulators of inhibitor of apoptosis (IAP) proteins.
Apoptosis inducing factor (AIF) and Endonuclease G, which translocate to the nucleus and cleave chromatin causing DNA fragmentation.
Easy Detection of the Mitochondrial Permeability Transition Event
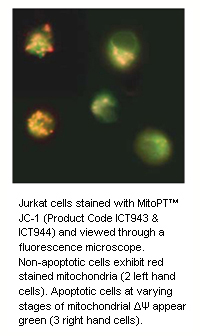
During apoptosis, the electrochemical gradient across the mitochondrial membrane collapses. This is called the mitochondrial permeability transition (PT) event. It is mediated by the opening of PT pore complexes comprised of adenine nucleoside translocator (ANT) in the inner mitochondrial membrane interacting with voltage dependent anion channel (VDAC) in the outer mitochondrial membrane. Cytosolic components along with osmotic water diffuse through the open channel into the mitochondrial matrix. This eventually swells and ruptures the mitochondrion releasing pro-apoptotic proteins, e.g. cytochrome c, into the cytosol.
Our quick and easy-to-use JC-1, TMRE and TMRM MitoPT™ Kits allow clear differentiation between non-apoptotic and apoptotic cells through the simple and reproducible fluorescence detection of the mitochondrial permeability transition event.
Our MitoPT™ Kit Range
More quality apoptosis antibodies and kits product ranges
References
Mitochondrial Released Proteins
-
Antonsson, B. et al. (2000) Bax oligomerization is required for channel-forming activity in liposomes and to trigger cytochrome c release from mitochondria.
Biochem J. 345:271-278. -
Cecconi, F. et al. (1998) Apaf1 (CED-4 homolog) regulates programmed cell death in mammalian development.
Cell 94:727-37. -
Desagher, S. et al. (1999) Bid-induced Conformational Change of Bax Is Responsible for Mitochondrial Cytochrome c Release during Apoptosis.
J. Cell Biol. 144:891-901. -
Hu, Y. et al. (1998) Bcl-XL interacts with Apaf1 and inhibits Apaf1-dependent caspase-9 activation.
PNAS 95:4386-91. -
Kroemer, G. and Reed, J.C. (2000) Mitochondrial control of cell death.
Nature Med. 6(5):513-519. -
Li, L.Y. et al. (2001) Endonuclease G is an apoptotic DNase when released from mitochondria.
Nature. 412:95-99. -
Li, P. et al. (1997) Cytochrome c and dATP-dependent formation of Apaf-1/caspase-9 complex initiates an apoptosis protease cascade.
Cell 91:479-489. -
Luo, X. et al. (1998) Bid, a Bcl2 interacting protein, mediates cytochrome c release from mitochondria in response to activation of cell surface death receptors.
Cell 94:481-490. -
Neame, S.J. et al. (1998) Blocking cytochrome c activity within intact neurons inhibits apoptosis.
J. Cell Biol. 142:1583-1593. -
Pan, G. et al. (1998) Caspase-9, Bcl-XL and Apaf-1 form a ternary complex.
J. Biol. Chem. 273:5841-5845. -
Ricci, J. et al. (2003) Caspase-mediated loss of mitochondrial function and generation of reactive oxygen species during apoptosis.
J. Cell Biol. 160 (1):65-75. -
Russell, J.W. et al. (2002) High glucose induced oxidative stress and mitochondrial dysfunction in neurons.
FASEB J. 16:1738-1748. -
Strauss, K. M. et al. (2005) Loss of function mutations in the gene encoding Omi/HtrA2 in Parkinson’s disease.
Human Molecular Genetics 14:2099-2111. -
Susin, S.A. et al. (1999) Molecular characterization of mitochondrial apoptosis - inducing factor.
Nature. 397:441-6. -
Suzuki, Y. et al. (2001) A Serine Protease, HtrA2, is released from the mitochondria and interacts with XIAP, inducing cell death.
Mol. Cell. 8:613-621. -
Verhagen, A. M. et al. (2001) Inhibitor of apoptosis proteins and their relatives. IAP’s and other BIRPS’s.
Genome Biol. 2(7):Reviews 3009. -
Wlodkowic, D. et al. (2006) Multiparametric analysis of HA14-1-induced apoptosis in follicular lymphoma cells.
Leukemia Res. 30:1187-1192. -
Wu, G. et al. (2000) Structural basis of IAP recognition by Smac/DIABLO.
Nature. 408:1008-12. -
Zou, H. et al. (1997) Apaf1, a human protein homologous to C. elegans CED-4, participates in cytochrome c-dependent activation of caspase-3.
Cell 90:405-13.
Mitochondrial Permeability Transition Event
-
Cossarizza, A. et al. (1993) A new method for the cytofluorimetric analysis of mitochondrial membrane potential using the J-aggregate forming lipophilic cation 5,5’,6,6’-tetrachloro-1,1’,3,3’-tetraethylbenzimidazolcarbocyanine iodide (JC-1) Biochem.
Biophys. Res. Commun. 197:40-45. -
Ehrenberg, B. V. et al. (1988) Membrane potential can be determined in individual cells from the nernstian distribution of cationic dyes.
Biophys. J. 53:785-794. -
Farkas, D. L. et al. (1989) Simultaneous imaging of cell and mitochondrial membrane potentials.
Biophys. J. 56:1053-1069. -
Huang, S. (2002) Development of a high throughput screening assay for mitochondrial membrane potential in living cells.
J. Biomol. Screen. 7(4):383-389. -
Lim, M. L. R. et al. (2001) The protonophore CCCP induces mitochondrial permeability transition without cytochrome c release in human osteosarcoma cells.
FASEB J . 503:69-74. -
Reers, M. et al. (1991) J-aggregate formation of a carbocyanine as a quantitative fluorescent indicator of membrane potential.
Biochemistry 30:4480-4486. -
Smiley, S. T. et al. (1991) Intracellular heterogeneity in mitochondrial membrane potentials revealed by a J-aggregate-forming lipophilic cation JC-1.
Proc. Natl. Acad. Sci. 88:3671-3675. -
Wong, A. and Cortopassi, G. A. (2002) High-throughput measurement of mitochondrial membrane potential in a neural cell line using a fluorescence plate reader.
Biochem. Biophys. Res. Comm. 29 8:750-754.




