Affinity Maturation of HuCAL® Antibodies
-
Monoclonal Generation
-
Custom Recombinant Monoclonal Antibody Generation
-
HuCAL® Antibody Generation Process
- Antigens and antigen expression services
- Guided Selection Strategies
- Affinity maturation
- Selection of antibodies based on binding strength
- Extended QC services
- Fab antibody formats and epitope tags
- Conversion of Fab to full immunoglobulin format
- Fab Antibody Production
- Screening and Pair Selection using the Bio-Plex® System
- Flow cytometry antibody screening
-
HuCAL® Antibody Generation Process
-
Custom Recombinant Monoclonal Antibody Generation
s
Simplified Sourcing via Scientist.com and ScienceExchange

Access Bio-Rad custom antibody generation services through the Scientist.com and ScienceExchange marketplaces.
s
Custom Antibody Project Inquiry Form
A personal, no obligation quotation for a custom monoclonal antibody generation project
s
My Green Lab Certified!

Our custom antibody services based in Neuried (Munich), Germany, are My Green Lab certified!
s
Contact our Custom Antibody Specialists
Tel: +49 (0) 89 80 90 95 45
Fax: +49 (0) 89 80 90 95 50
Office: Bio-Rad AbD Serotec GmbH, Campus Neuried, Anna-Sigmund-Str. 5, 82061 Neuried, Germany
Improving the binding strength of recombinant monoclonal antibodies
In some cases sensitivity of an assay can be improved by incorporating antibodies with a high affinity for the target antigen; for example, when measuring an antigen known to be at a low concentration in the sample or when using anti-peptide antibodies for immunoaffinity capture and enrichment.
The affinity of recombinant monoclonal antibodies generated using HuCAL technology is comparable to monoclonal antibodies made using conventional animal methods. For protein antigens it is generally in the range 1-100 nM, and in many cases sub-nanomolar affinities are achieved.
In contrast to animal-derived antibodies from serum or hybridoma supernatant, HuCAL antibodies can be easily optimized for affinity, since the antibody DNA sequence is known and available. The specificity of the antibody usually remains unchanged during the antibody maturation process. The affinity improvement will depend on the affinity of the parental binder and the nature of the antigen. Using our methods, improvement of at least 10-fold is routinely achieved and improvements of more than 1000-fold have also been reached.
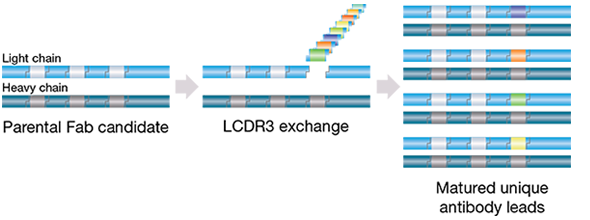
Affinity maturation is an optional step that can be performed once antigen-specific HuCAL antibodies have been selected. Binding characteristics are optimized by inserting pre-built CDR library cassettes (usually at LCDR3 or HCDR2), which have been diversified according to the natural repertoire of CDR sequences at unique flanking restriction sites, Figure 1.
Fig. 1. Optimization of binding characteristics of a HuCAL antibody occurs via CDR library creation for specific binders in a simple cloning step.
Affinity maturation of a single binder
This approach starts with an antibody selected from the standard antibody generation process that shows a very high specificity, but lacks sensitivity in the desired application. The LCDR3 or HCDR2 region of the parental antibody sequence is exchanged with highly diversified cassettes to generate a new antibody library of up to 108 antibodies differing only in the sequence of the exchanged CDR.
Phage panning on the antigen is repeated with higher stringency; for example, by increasing the number and/or length of the washing steps, or reducing the amount of antigen in the panning rounds. The result is affinity matured unique antibodies.
As part of the screening step, a high throughput off-rate determination is recommended for a minimum of 95 antibodies in order to select those with the highest affinities. These antibodies are then purified and the affinity is measured. The final improvement in affinity will depend on the affinity of the parental binder and the nature of the antigen.
A single binder affinity maturation project is likely to take 6-7 months. This includes the initial 8 weeks for Fab antibody generation, time to test antibodies and select the parental clone for maturation, a further 16 weeks for the generation of the new antibody libraries, panning on the antigen, koff-rate ranking, purification and final affinity determination.
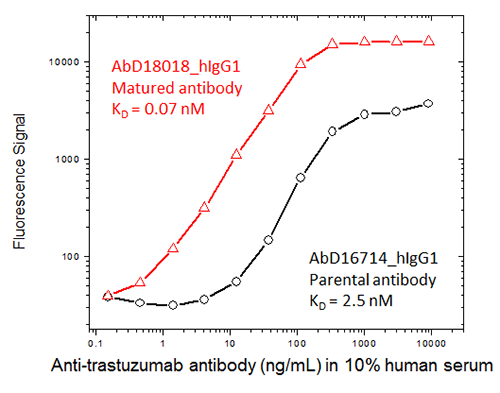
Fig. 2. An anti-trastuzumab antibody ranked as having the highest affinity after maturation (red) was compared with the non-matured, parental antibody (black) in an ADA bridging ELISA format assay; the data highlights the difference in sensitivity obtained with the affinity matured antibody.
Rapid affinity maturation
A more time efficient approach for affinity maturation is rapid pool maturation, RapMAT®. This method is recommended when it is known at the outset that a maturation step is likely to be required to meet very high affinity requirements for the antibodies.
Affinity maturation is incorporated as part of the standard HuCAL antibody generation process. Starting with the complete HuCAL antibody library of 45 billion Fab antibodies, two rounds of panning on the antigen are carried out. Following this, the pre-enriched pool of phage displaying antibodies specific to the target antigen is taken and for each antibody sequence the LCDR3 or HCDR2 is replaced in a pool cloning step to create a new library. Subsequently this library is used for two further rounds of panning applying a higher stringency. This RapMAT process takes approximately 16 weeks, Figure 3.
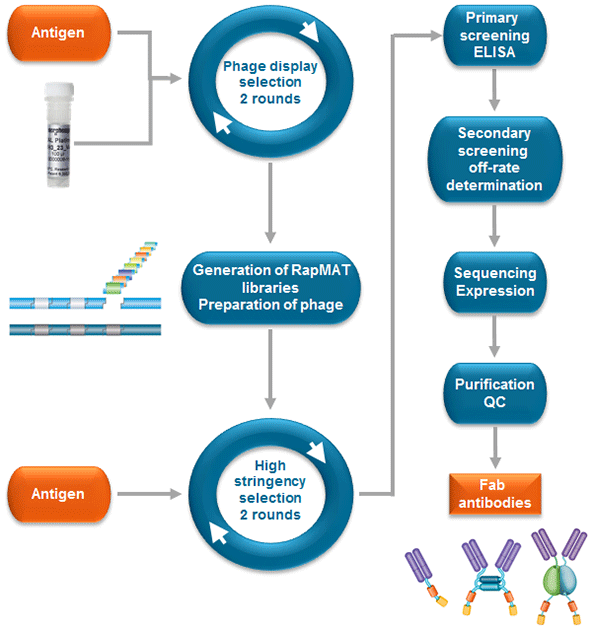
Fig. 3. Process flow for RapMAT rapid pool affinity maturation.
Our technical experts are looking forward to designing a strategy to generate the antibodies required for your assays and will be able to recommend an affinity maturation approach appropriate to your project needs.
References
-
Harth S et al. (2019). Generation by phage display and characterization of drug-target complex-specific antibodies for pharmacokinetic analysis of biotherapeutics.
mAbs 11:178-90 -
Ylera, F. et al. (2013). Off-rate screening for selection of high-affinity anti-drug antibodies.
Anal Biochem. 441:208-13 -
Steidl, S. et al. (2008). In vitro affinity maturation of human GM-CSF antibodies by targeted CDR-diversification.
Mol Immunol. 46:135-44