
Cell Activation Protocols
T cell stimulation for flow cytometry analysis
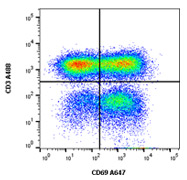
Fig. 1. PBMCs extracted from human blood were stimulated with PHA and LPS for 48 hrs. Activated T cells are seen by upregulated CD69 (MCA2806A647) expression on the CD3 (MCA463A488) positive population. Cells were gated on lymphocytes in the presence of Human Seroblock (BUF070A). Data acquired on the ZE5™ Cell Analyzer.
Activation and proliferation protocols provide an effective method to determine immunocompetence and cell reactivity. However choosing the right stimulation protocol and knowing when to use it may vary depending on what cell type you are starting with and the result you need.
To help you we have grouped a few simple protocols for T cells, with some background information so you can choose the right method. These protocols are useful when you need to measure activation markers (Figure 1), cytokine release, receptor expression and proliferation responses (Figure 2). It can also be used as a positive control for your experiment. The length of time in culture prior to analysis will determine what can be effectively measured. Activation markers are visible after a few hours whereas proliferation can take up to five days. Although these protocols will allow you to assess reactivity they will not give you information about specific effector function nor allow you to identify specific T cell subsets.
FC15 Unprimed T Cell Activation – Pharmacological Method
This protocol provides a general method to activate unprimed T cells using non-specific agents such as Phorbol 12-myristate 13-acetate (PMA) in combination with ionomycin. This protocol is ideal when maximal stimulation that is not reliant on specific cell receptors is required. Note that some activation reagents will require the presence of accessory cells such as antigen presenting cells in the culture, which may need LPS activation in addition to the T cells to function effectively. You may have to irradiate accessory cells to prevent interference into the T cell population.
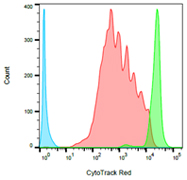
Fig. 2. Human PBLs were stained with CytoTrack Red 628/643 Cell Proliferation Assay Kit (1351205) and stimulated for five days with PHA. Cells that have proliferated show a reduction in the amount of dye with each cell division. Six peaks corresponding to each division are visible in this experiment. You can see stimulated cells (in red), labeled cells but unstimulated (in green) and unlabelled cells (in blue). Data acquired on the ZE5 Cell Analyzer.
FC16 Unprimed T Cell Activation - Antibody Stimulation Method
This protocol provides a general method for using anti-CD3 and anti-CD28 antibodies when the anti-CD3 antibody is bound to a plate. It is possible to use anti-CD3 in solution, however, the activation is dependent on receptor cross-linking, so antigen presenting cells or a secondary antibody may be required. If you are using soluble antibodies, you will need a low endotoxin format of the antibody to avoid the effects of preservatives and stabilizers on your cultures. Some clones of anti-CD3 and anti-CD28 have been shown to be less effective at cell activation than others. If you want to stain for CD3 or CD28 post activation, clones with alternative binding sites may be required as the binding sites may be blocked.
FC17 Primed T Cell Activation - Antigen Presenting Cell Co-culture
This protocol provides a general method for activating primed T cells in a more physiological manner compared to the two previous protocols (FC15 and FC16). This protocol requires the preparation of antigen presenting cells such as dendritic cells prior to harvesting T cells and an appropriate source of antigen.
FC18 Measuring Proliferation using Cell Permeable Dyes
Cell proliferation can be measured in several ways such as basic counting of cells and H3 thymidine uptake. Here we describe a simple protocol, requiring no radioactivity, to label cells with cell permeable CytoTrack™ Dyes. These dyes are available in a range of excitation and emission wavelengths to easily fit into multicolor flow cytometry panels.
Troubleshooting
There are many factors that can influence the effectiveness of these protocols and they should be tested empirically to determine the optimal conditions in your laboratory. The concentration of reagents and cells, the culture and labeling time, culture conditions, source of serum and type of tissue culture plate, and the source of cells, such as mouse strain, can all have an effect on the responsiveness of your T cells in culture and may need to be optimized.
As with all protocols, including the right controls is important. Negative controls, such as unstimulated or unlabeled cells, will enable you to distinguish activated from non-activated cells. In addition, the inclusion of viability dyes is essential to remove dead cells from your analysis, which may give false positives. Further information on controls in flow cytometry and common flow cytometry protocols are available. You can also view our database of over 3,000 flow cytometry validated antibodies, kits and supporting reagents.
Although the protocols described here have been designed for flow cytometry analysis, they are suitable for use in other applications such as ELISA, where cytokine output could be measured and microscopy, to look at receptor capping, with the appropriate modifications. For further information on these techniques, visit our dedicated antibody application pages.