ELISA: Sandwich Antibody Pair Generation

- On This Page
- Strategies for selecting antibody pairs
- Timelines
s
Simplified Sourcing via Scientist.com and ScienceExchange

Access Bio-Rad custom antibody generation services through the Scientist.com and ScienceExchange marketplaces.
s
Custom Antibody Project Inquiry Form
A personal, no obligation quotation for a custom monoclonal antibody generation project
s
My Green Lab Certified!

Our custom antibody services based in Neuried (Munich), Germany, are My Green Lab certified!
s
Contact our Custom Antibody Specialists
Tel: +49 (0) 89 80 90 95 45
Fax: +49 (0) 89 80 90 95 50
Office: Bio-Rad AbD Serotec GmbH, Campus Neuried, Anna-Sigmund-Str. 5, 82061 Neuried, Germany
Strategies for selecting antibody pairs
Selection of antibodies using HuCAL® technology is performed in vitro. This enables greater flexibility for antibody generation than is possible using traditional methods based on the immunization of animals.
‘Guided Selection’ is the term we use for all selection strategies used for the identification of antibodies to specific epitopes or epitope areas. Two strategies are typically used for the generation of sandwich pair antibodies.
- Antibody selection (panning) against the antigen is followed by testing all possible combinations of the selected antibodies as capture and detection antibodies in a sandwich immunoassay (ELISA microtiter plate-based method or bead-based assay using Bio-Plex® 3D Suspension Array System). A final titration ELISA is performed on the five best matched pairs and a report is generated to allow the client to select the desired antibodies. The best combination of capture and detection antibodies is identified by this method. This method depends on the number of antibodies selected during the panning of the antibody library and their properties, which in turn depends on the antigen used.
- If technique 1 is unsuccessful a second method is used. Standard panning against the antigen is followed by ELISA screening of the purified antibodies for the best capture antibody. Labeled antigen, e.g. biotinylated, is required for detection. The best capture antibody is then used to capture the antigen and a subsequent panning is performed on the antibody-antigen complex (Figure 1). The capture antibody is used to pre-clear the library and an isotype control antibody is used for blocking, ensuring depletion of capture-antibody specific antibodies. Selected antibodies will bind the captured antigen and therefore are matched detection antibodies.
More about high-throughput pair identification using the Bio-Plex Multiplex Immunoassay System.
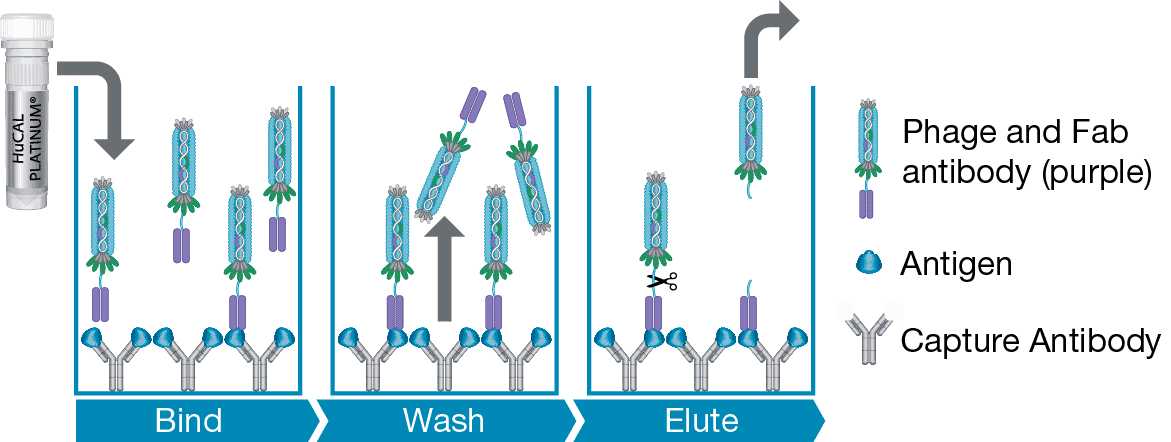
Fig. 1. Selection with a captured antigen. The antigen is captured by an antibody, previously isolated from the library. Panning is carried out on the complex to select for the detection antibody.
Timelines
The standard panning project (strategy 1) generates Fab antibodies in as little as 8 weeks. A further 1-2 weeks is required to find the best combination of capture and detection antibodies. The second strategy requires an initial panning of the library to select the capture antibody (8 weeks, plus 1-2 weeks testing to find the best antibody) and a second panning to select the detection antibody (8 weeks), plus 1-2 weeks for testing to find the best antibody pair.
Depending on the availability of your target of interest, the project can also start with the sequence of your protein and include an antigen generation step, which can be done using gene synthesis and expression or peptide synthesis and coupling. An additional 5-10 weeks is needed for this depending on the route chosen.
Three steps from gene to highly specific antibody pairs ready for assay development
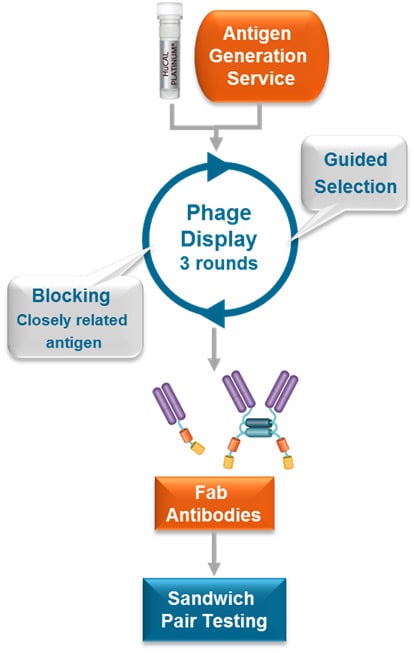
Step 1: Antigen generation
- Gene synthesis and expression
- Peptide synthesis and coupling
Step 2: Antibody generation
- Primary screening and sequencing
- Purification and labeling
Step 3: Sandwich ELISA testing
- Sandwich ELISA screen, all against all
- Final titration ELISA
Fig. 2. Process flow for sandwich ELISA antibody pair selection using HuCAL technology.





