BrdU Staining Experiments: Tips for Success
At Bio-Rad, we have developed a set of tips for your BrdU labeling experiments to ensure robust data generation.
- Select the right BrdU antibody for your experiment.
- Know your anti-BrdU antibody cross-reactivity.
- Keep your BrdU fresh.
- Perform a BrdU titration.
- Optimize your DNA denaturation step.
- Optimize your washing steps.
- Carefully select your secondary antibody.
- Include suitable controls.
- Take your tissue sample size into account when performing immunohistochemistry (IHC) experiments.
- Consider spectral overlap in multichannel flow cytometry.
Select the Right BrdU Antibody for Your Experiment
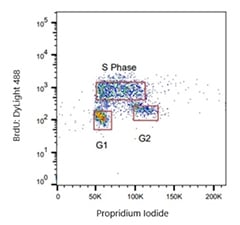
Fig. 1. Jurkat cells were treated with 100 μM BrdU and cells were stained with clone RF04-2 Rat Anti-BrdU Antibody (MCA6143). As a secondary antibody, Rabbit F(ab')2 Anti-Rat IgG: DyLight488 Conjugated Antibody (STAR16D488GA) was used. Cells actively synthesizing DNA (S Phase) show BrdU incorporation, resulting in a bright signal from BrdU:DyLight488.
Antibody selection is one of the most important considerations when planning an experiment. Ideally, your chosen antibody should have supporting data that shows characteristic positive staining of BrdU, in your chosen application. This can usually be found on the datasheet or product webpage.
For example, if you are analyzing BrdU incorporation via flow cytometry, you should look for supplier data showing the characteristic “hook profile” when the nuclear dye is plotted on the X-axis and BrdU is plotted on the Y-axis. Using this profile, you should be able to discriminate between G1, S, and G2 phases. The cells that are in the S-phase of the cell cycle have brighter staining than those in G1 or G2 phase (Figure 1).
In addition, choose an anti-BrdU antibody with a different host species to the cells or tissue you plan to use for your experiment. For the best lot-to-lot consistency and for long-term security of supply, use non-animal derived, recombinant anti-BrdU antibodies. You should also perform titration experiments in order to determine the optimal concentration of your anti-BrdU antibody.
Know Your Anti-BrdU Antibody Cross-Reactivity
BrdU and thymidine are structurally similar. Their structural difference is illustrated in Figure 2; in the case of BrdU, the 5-methyl group (shown in blue) of thymidine has been substituted by bromine (highlighted in green) (Kolb et al. 1999, Sivakumar et al. 2004). As an alternative to bromine, the 5-methyl group of thymidine can be replaced with other halogens such as chlorine (5′-chloro-2′-deoxyuridine, CldU) and iodine (5′-iodo-2′-deoxyuridine, IdU) (Sivakumar et al. 2004), to create different thymidine analogs.
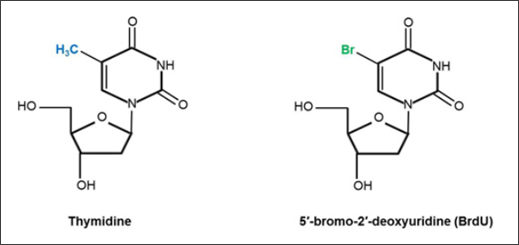
Fig. 2. Chemical structures of the DNA nucleoside thymidine and its analog BrdU. Structures adapted from Liboska et al. (2012) (BrdU) and Young et al. (1969) (thymidine).
Anti-BrdU antibodies are likely to recognize other thymidine analogs due to their structural similarities. If you are looking to incorporate different analogs at different time points, for example when performing a dual pulse labeling experiment, check if your BrdU antibody has been tested for cross-reactivity with thymidine analogs.
It is also useful to look for data demonstrating a lack of cross-reactivity with thymidine. If your anti-BrdU antibody cross-reacts with thymidine in the DNA of cells, this could result in a false positive signal.
Keep Your BrdU Fresh
Add freshly prepared BrdU to appropriate growth media for your cells or tissue. Remember that the half-life of BrdU at 4°C is short, so make sure your BrdU stock solution is stored at -20°C.
Perform a BrdU Titration
The ideal concentration of BrdU and incubation time for optimal labeling depends on the rate of cell division, and needs to be optimized for your cells. Perform titration experiments to determine the concentration of BrdU that will give the best level of incorporation without inducing cytotoxicity.
Optimize Your DNA Denaturation Step
For successful staining, it is important to include a DNA denaturation step to allow the antibody access to the incorporated BrdU. One common approach uses depurination and cleavage of DNA by acids such as hydrochloric acid. Other treatments with copper ions, heat, nucleases, and sodium hydroxide have also been reported (Liboska et al. 2012, Kennedy et al. 2000). Optimize your denaturation step by establishing the hydrochloric acid concentration, temperature, and incubation period that give the best BrdU detection and keep the protein structure in the cells intact.
Optimize Your Washing Steps
After the denaturation step, ensure you perform sufficient washing in an appropriate amount of wash buffer. Any remaining acid that is not washed away could denature your antibody and affect the ability to detect BrdU. Keep in mind that the wash steps also need to be optimized for the anti-BrdU antibody you are using to give the best signal-to-background ratio.
Carefully Select Your Secondary Antibody
If you are planning to detect your anti-BrdU antibody with a secondary antibody, rather than a directly conjugated anti-BrdU antibody, background staining may be reduced by using a secondary antibody raised in a different species to the target cells/tissues.
Include Suitable Controls
Proper controls add meaning to your data. You should make sure that all the relevant controls are included. These could be:
Negative controls
To observe any effect of the solvent on your cells, treat a sample of cells with only the solvent that the stock BrdU solution is diluted in. For example, if your stock BrdU solution is diluted in dimethyl sulfoxide (DMSO), you should prepare negative control cells treated with DMSO only. It is essential that the volume of solvent administered to your negative control group is the same as the volume of BrdU in DMSO administered to your treatment group, and that the incubation period is identical for both samples.
Isotype controls
You should also consider isotype controls. This involves staining a sample of your cells or tissue with an antibody that binds to an unrelated target (not designed to target BrdU) and of the same isotype as the antibody targeting BrdU.
Secondary only antibody controls
If you are using secondary antibodies, it is advisable to run samples that are assayed with just the secondary antibody (without using the primary BrdU antibody), to check for background staining that results from nonspecific binding of the secondary antibody.
Take Your Tissue Sample Size into Account While Performing IHC Experiments
BrdU staining of tissue samples can be detected using immunohistochemistry. Note, the antibodies detecting BrdU need to penetrate all cells within a fixed tissue, so the size of your tissue sample will affect BrdU staining. You can overcome this by sectioning the tissue or by prolonging antibody incubation (Salic and Mitchison 2008).
Consider Spectral Overlap in Multichannel Flow Cytometry
If you use flow cytometry to analyze BrdU incorporation, remember to select a nuclear dye that has no spectral overlap with the fluorophore chosen for the BrdU antibody. One simple way to check this is to use a spectraviewer tool.
Visit bio-rad-antibodies.com/fluorescent-spectraviewer.html to access Bio-Rad’s free spectraviewer tool.
References:
- Liboska et al. (2012). Most anti-BrdU antibodies react with 2'-deoxy-5-ethynyluridine -- the method for the effective suppression of this cross-reactivity. PLOS One, 7 e51679.
- Kennedy BK et al. (2000). Nuclear organization of DNA replication in primary mammalian cells. Genes Dev 15, 2,855–2,868.
- Kolb B et al. (1999). Embryonic and postnatal injections of bromodeoxyuridine produce age-dependent morphological and behavioral abnormalities. J Neuro 19, 2,337–2,346.
- Liboska R et al. (2012). Most anti-BrdU antibodies react with 2′-deoxy-5′-ethynyluridine – the method for the effective suppression of this cross-reactivity. PLoS One 7, e51679.
- Salic A and Mitchison J (2008). A chemical method for fast and sensitive detection of DNA synthesis in vivo. PNAS 7, 2,415–2,420.
- Sivakumar S et al. (2004). In vivo labeling of fission yeast DNA with thymidine and thymidine analogs. Methods 33, 213–219.
- Young DW et al. (1969). The crystal and molecular structure of thymidine. Aca Cryst B25, 1,423–1,432.







