5-Methylcytidine antibody | 33D3









Mouse anti 5-Methylcytidine
- Product Type
- Monoclonal Antibody
- Clone
- 33D3
- Isotype
- IgG1
- Specificity
- 5-Methylcytidine
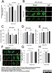
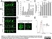
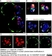
| Mouse anti 5-methylcytidine antibody, clone 33D3 recognizes 5-methylcytidine, a modified base found in the DNA of plants and vertebrates. Methylation of DNA is an epigenetic process that stably alters the expression of genes in cells as they divide and differentiate into specific tissues. The resulting change is normally permanent and unidirectional, preventing a cell from reverting to a stem cell or converting into another type of tissue. In cancer biology, DNA hypermethylation is associated with gene silencing while hypomethylation is reported to be associated with disease progression (Sincic & Herceg, 2011). Mouse anti 5-MeCyd antibody, clone 33D3 is specific to the methylated base and has minimal reactivity to non-methylated cytidine or cytosine (Reynaud et al.1992) Mouse anti 5-MeCyd antibody, clone 33D3 has been reported for use in methylated DNA immunoprecipitation (MeDIP) (Pontes et al. 2007). |
- Target Species
- Broad
- Species Cross-Reactivity
-
Target Species Cross Reactivity Human Rat Mouse Bovine - N.B. Antibody reactivity and working conditions may vary between species.
- Product Form
- Purified
- Preparation
- Purified IgG prepared by affinity chromatography on Protein A from tissue culture supernatant
- Buffer Solution
- Phosphate buffered saline
- Preservative Stabilisers
- 0.09% Sodium Azide (NaN3)
- Approx. Protein Concentrations
- IgG concentration 1.0 mg/ml
- Fusion Partners
- Spleen cells from immunised Balb/c mice were fused with cells of the Sp2/0-Ag 14 myeloma cell line.
- Regulatory
- For research purposes only
- Guarantee
- 12 months from date of despatch
Avoid repeated freezing and thawing as this may denature the antibody. Storage in frost-free freezers is not recommended.
| Application Name | Verified | Min Dilution | Max Dilution |
|---|---|---|---|
| ELISA | |||
| Flow Cytometry 1 | |||
| Immunoblotting | |||
| Immunofluorescence | |||
| Immunohistology - Frozen | |||
| Immunohistology - Paraffin 2 | |||
| Methylated DNA Immunoprecipitation |
- 1Membrane permeabilization may be required prior to staining. Cell pretreatment before staining is described in Giraldo et al (2007)..
- 2This product requires antigen retrieval using heat treatment prior to staining of paraffin sections.Sodium citrate buffer pH 6.0 is recommended for this purpose.
- Flow Cytometry
- Use 10μl of the suggested working dilution to label 1x106 cells in 100μl
| Description | Product Code | Applications | Pack Size | List Price | Your Price | Quantity | |
|---|---|---|---|---|---|---|---|
| Rabbit F(ab')2 anti Mouse IgG:RPE | STAR12A | F | 1 ml |
|
Log in | ||
| List Price | Your Price | ||||||
|
|
Log in | ||||||
| Description | Rabbit F(ab')2 anti Mouse IgG:RPE | ||||||
| Rabbit F(ab')2 anti Mouse IgG:HRP (Human Adsorbed) | STAR13B | C E P RE WB | 1 mg |
|
Log in | ||
| List Price | Your Price | ||||||
|
|
Log in | ||||||
| Description | Rabbit F(ab')2 anti Mouse IgG:HRP (Human Adsorbed) | ||||||
| Rabbit F(ab')2 anti Mouse IgG:FITC | STAR9B | F | 1 mg |
|
Log in | ||
| List Price | Your Price | ||||||
|
|
Log in | ||||||
| Description | Rabbit F(ab')2 anti Mouse IgG:FITC | ||||||
References for 5-Methylcytidine antibody
-
Reynaud, C. et al. (1992) Monitoring of urinary excretion of modified nucleosides in cancer patients using a set of six monoclonal antibodies.
Cancer Lett. 61 (3): 255-62. -
Habib, M. et al. (1999) DNA global hypomethylation in EBV-transformed interphase nuclei.
Exp Cell Res. 249 (1): 46-53. -
Hernandez-Blazquez, F.J. et al. (2000) Evaluation of global DNA hypomethylation in human colon cancer tissues by immunohistochemistry and image analysis.
Gut. 47 (5): 689-93. -
Giraldo, A.M. et al. (2007) DNA methylation and histone acetylation patterns in cultured bovine fibroblasts for nuclear transfer.
Mol Reprod Dev. 74 (12): 1514-24. -
Shen, R. et al. (2009) Reversibility of aberrant global DNA and estrogen receptor-alpha gene methylation distinguishes colorectal precancer from cancer.
Int J Clin Exp Pathol. 2 (1): 21-33. -
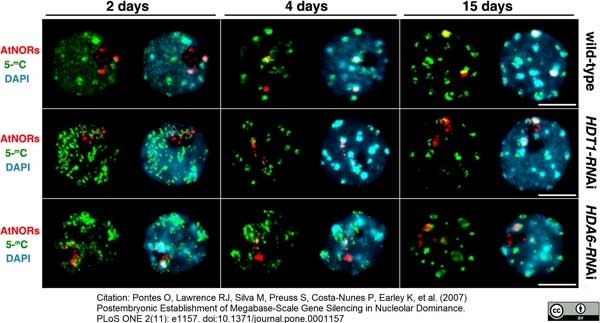
Pontes, O. et al. (2007) Postembryonic establishment of megabase-scale gene silencing in nucleolar dominance.
PLoS One. 2 (11): e1157. -
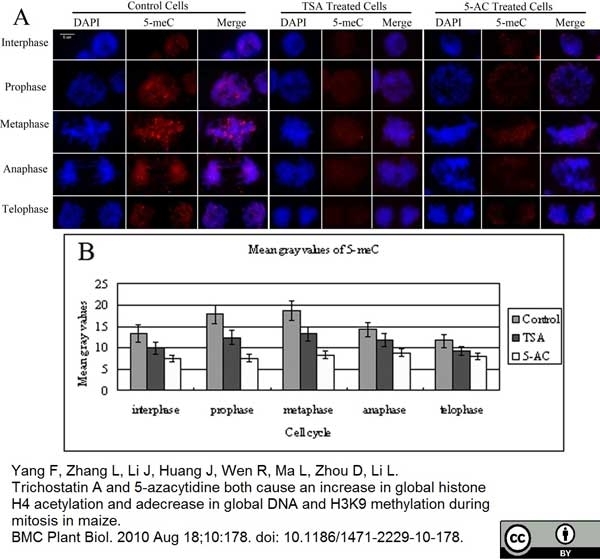
Yang, F. et al. (2010) Trichostatin A and 5-azacytidine both cause an increase in global histone H4 acetylation and a decrease in global DNA and H3K9 methylation during mitosis in maize.
BMC Plant Biol. 10: 178. -
Suter, J.D. et al. (2010) Label-free DNA methylation analysis using opto-fluidic ring resonators.
Biosens Bioelectron. 26: 1016-20.
View The Latest Product References
-
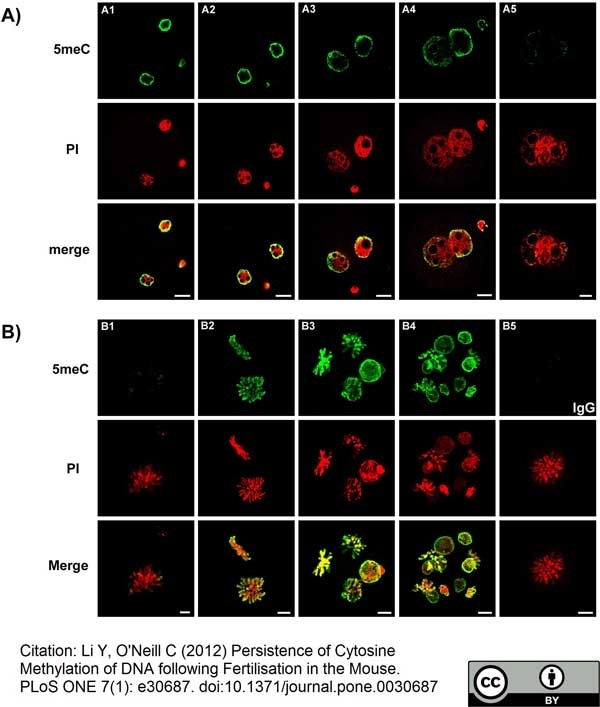
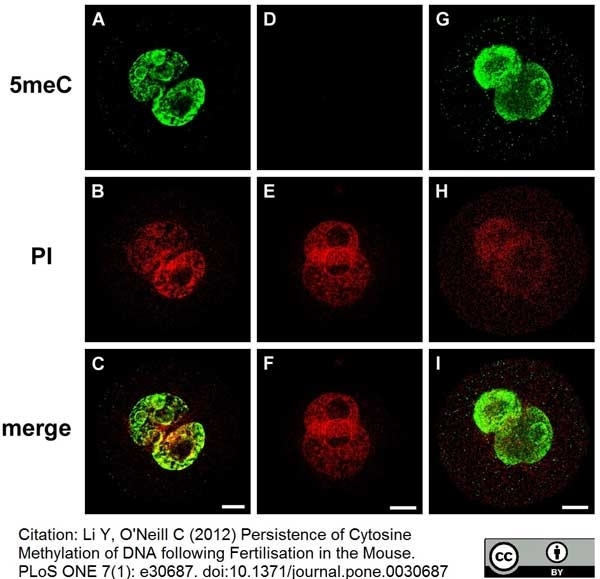
Li, Y. and O'Neill, C. (2012) Persistence of cytosine methylation of DNA following fertilisation in the mouse.
PLoS One. 7:e30687. -
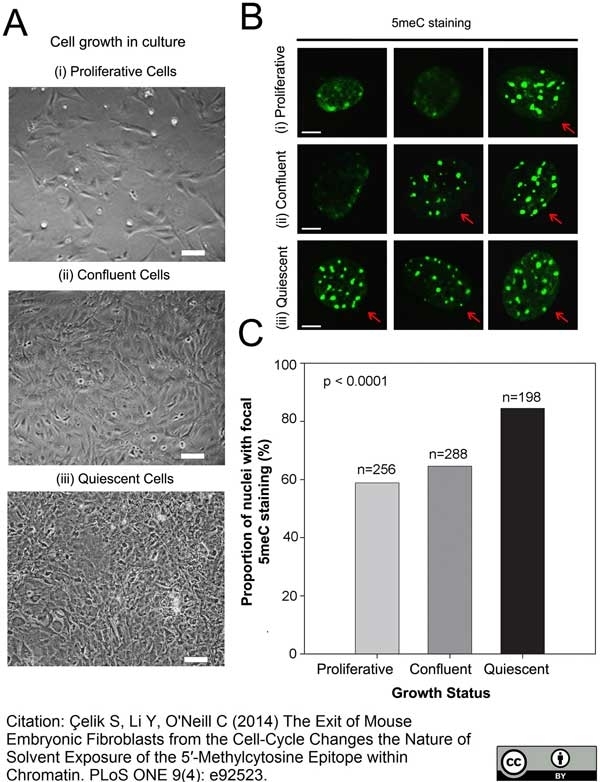
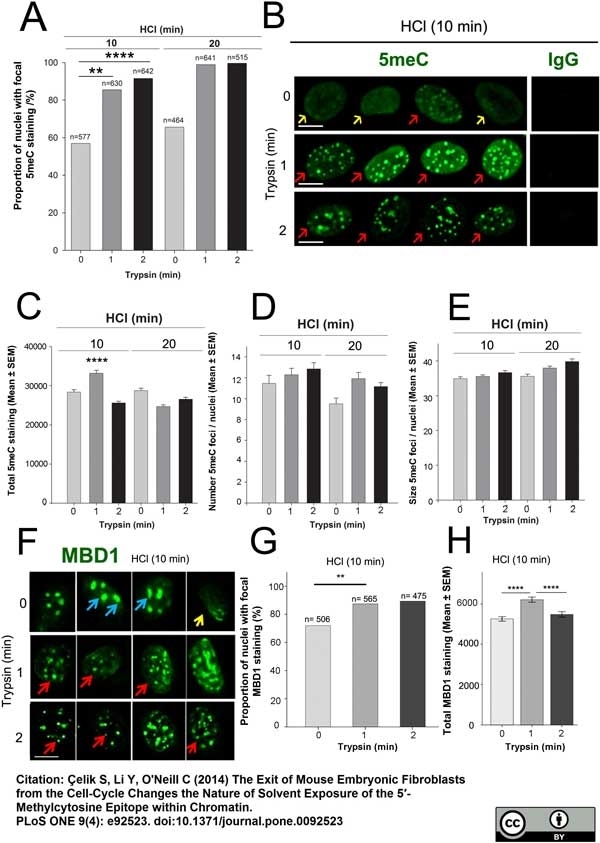
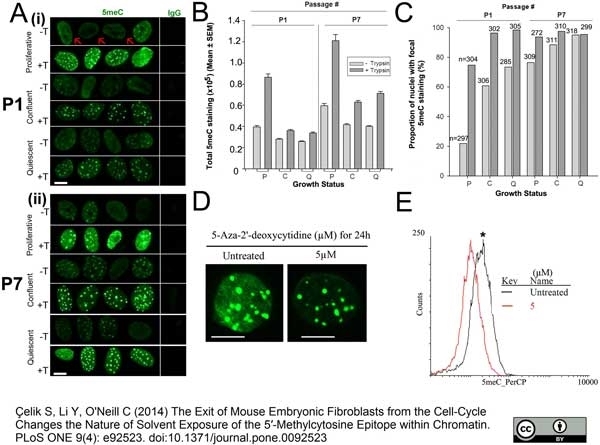
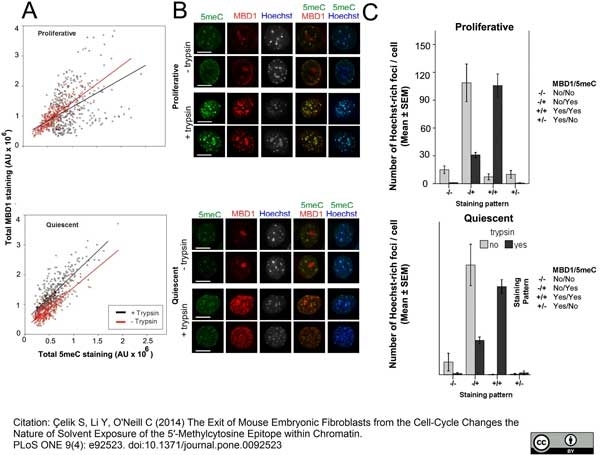
Çelik, S. et al. (2014) The Exit of Mouse Embryonic Fibroblasts from the Cell-Cycle Changes the Nature of Solvent Exposure of the 5'-Methylcytosine Epitope within Chromatin.
PLoS One. 9: e92523. -
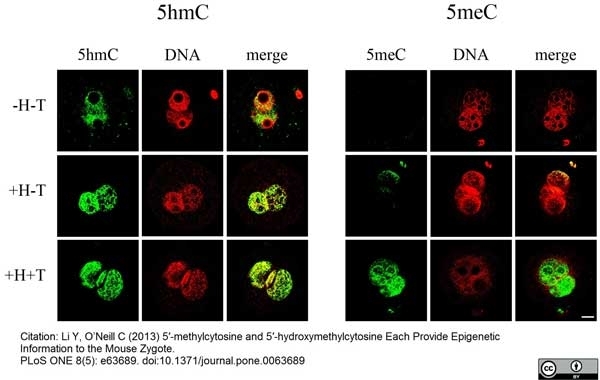
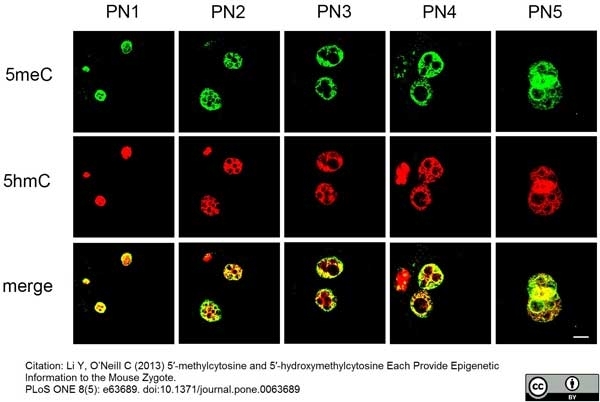
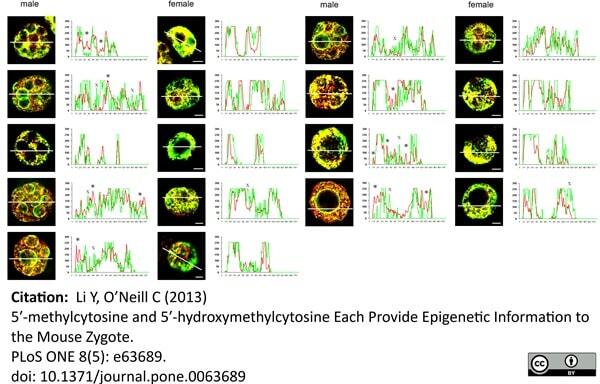
Li, Y. & O'Neill, C. (2013) 5'-Methylcytosine and 5'-hydroxymethylcytosine each provide epigenetic information to the mouse zygote.
PLoS One. 8 (5): e63689. -
Leter, G. et al. (2014) Exposure to perfluoroalkyl substances and sperm DNA global methylation in Arctic and European populations.
Environ Mol Mutagen. 55 (7): 591-600. -
Desjobert, C. et al. (2015) Combined analysis of DNA methylation and cell cycle in cancer cells.
Epigenetics. 10 (1): 82-91. -
Çelik-Uzuner S et al. (2016) Measurement of global DNA methylation levels by flow cytometry in mouse fibroblasts.
In Vitro Cell Dev Biol Anim. Aug 9. [Epub ahead of print] -
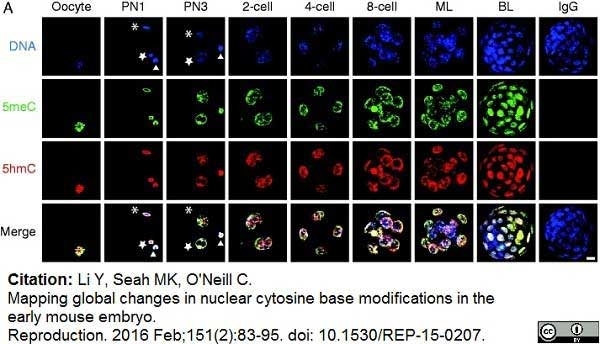
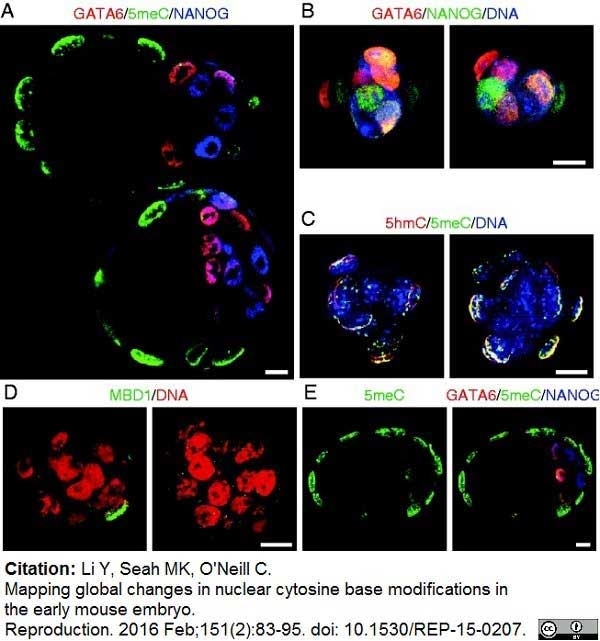
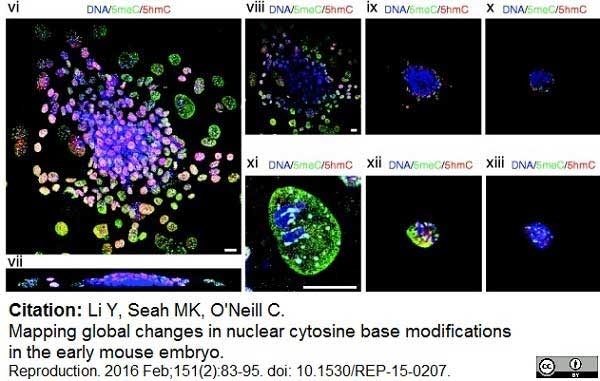
Li, Y. et al. (2016) Mapping global changes in nuclear cytosine base modifications in the early mouse embryo.
Reproduction. 151 (2): 83-95. -
Çelik-Uzuner, S. & O'Neill, C. (2021) Cellular Autofluorescence in Mouse Embryonic Fibroblasts Interferes with Antigen Detection Using Flow Cytometry.
J Fluoresc. Mar 27 [Epub ahead of print]. -
Daniel, S. et al. (2018) T cell epigenetic remodeling and accelerated epigenetic aging are linked to long-term immune alterations in childhood cancer survivors.
Clin Epigenetics. 10 (1): 138. -
Baixauli, F. et al. (2022) An LKB1-mitochondria axis controls T(H)17 effector function.
Nature. 610 (7932): 555-561.
Further Reading
-
Sinčić, N & Herceg, Z. (2011) DNA methylation and cancer: ghosts and angels above the genes.
Curr Opin Oncol. 23: 69-76.
- RRID
- AB_324056
MCA2201
If you cannot find the batch/lot you are looking for please contact our technical support team for assistance.
Please Note: All Products are "FOR RESEARCH PURPOSES ONLY"
View all Anti-Broad ProductsAlways be the first to know.
When we launch new products and resources to help you achieve more in the lab.
Yes, sign me up